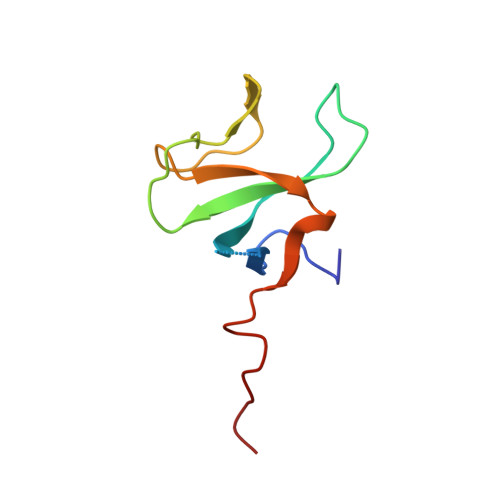
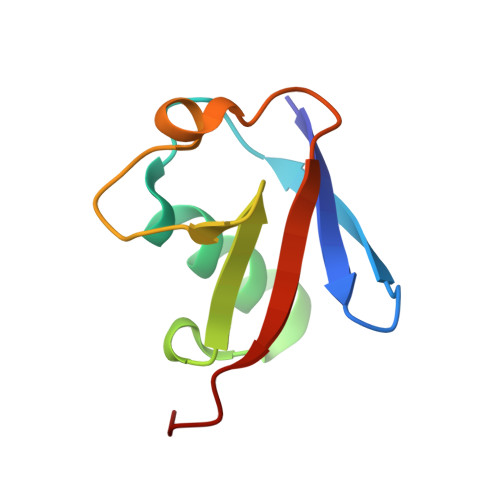
Regulation of CYLD activity and specificity by phosphorylation and ubiquitin-binding CAP-Gly domains.
Elliott, P.R., Leske, D., Wagstaff, J., Schlicher, L., Berridge, G., Maslen, S., Timmermann, F., Ma, B., Fischer, R., Freund, S.M.V., Komander, D., Gyrd-Hansen, M.(2021) Cell Rep 37: 109777-109777
- PubMed: 34610306 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2021.109777
- Primary Citation Related Structures:
7OWC, 7OWD - PubMed Abstract:
Non-degradative ubiquitin chains and phosphorylation events govern signaling responses by innate immune receptors. The deubiquitinase CYLD in complex with SPATA2 is recruited to receptor signaling complexes by the ubiquitin ligase LUBAC and regulates Met1- and Lys63-linked polyubiquitin and receptor signaling outcomes. Here, we investigate the molecular determinants of CYLD activity. We reveal that two CAP-Gly domains in CYLD are ubiquitin-binding domains and demonstrate a requirement of CAP-Gly3 for CYLD activity and regulation of immune receptor signaling. Moreover, we identify a phosphorylation switch outside of the catalytic USP domain, which activates CYLD toward Lys63-linked polyubiquitin. The phosphorylated residue Ser568 is a novel tumor necrosis factor (TNF)-regulated phosphorylation site in CYLD and works in concert with Ser418 to enable CYLD-mediated deubiquitination and immune receptor signaling. We propose that phosphorylated CYLD, together with SPATA2 and LUBAC, functions as a ubiquitin-editing complex that balances Lys63- and Met1-linked polyubiquitin at receptor signaling complexes to promote LUBAC signaling.
- Division of Protein and Nucleic Acid Chemistry, MRC Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge CB2 0QH, UK; Department of Biochemistry, University of Oxford, South Parks Road, Oxford OX1 3QU, UK. Electronic address: paul.elliott@bioch.ox.ac.uk.
Organizational Affiliation: