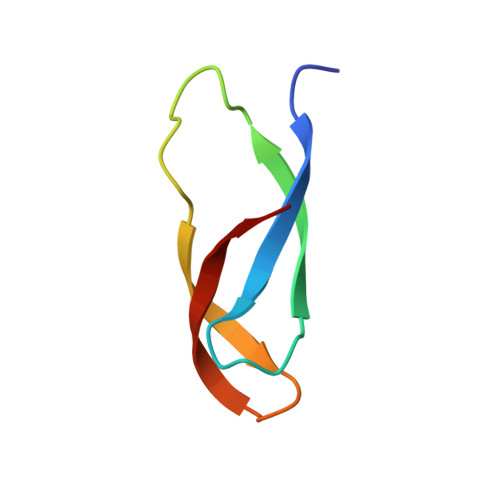
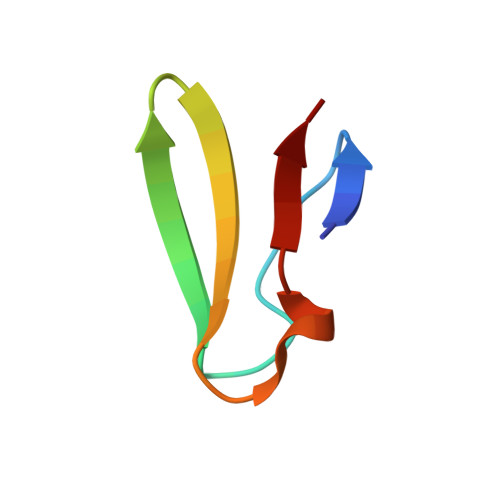
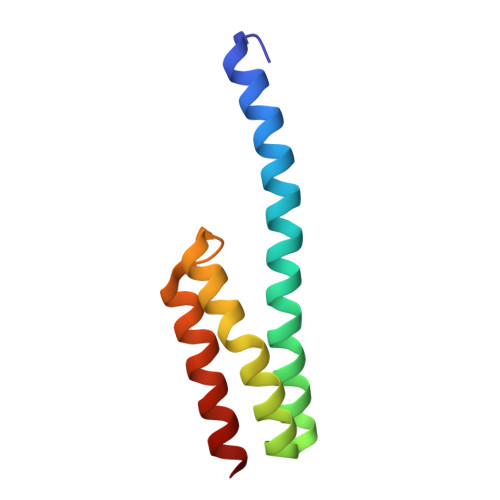
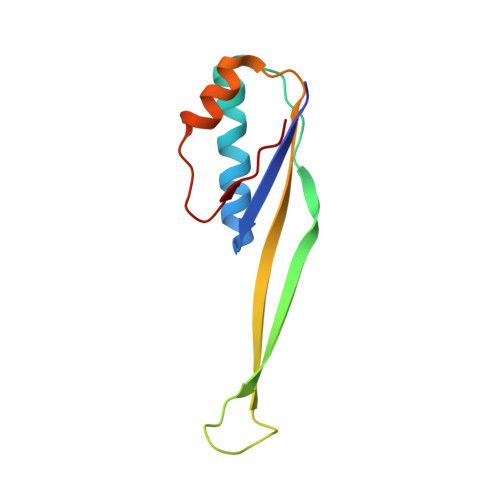
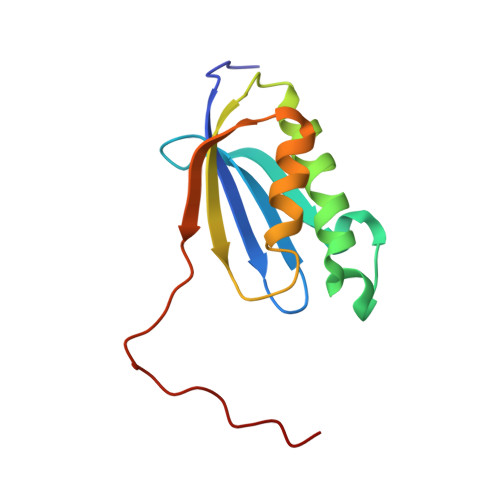
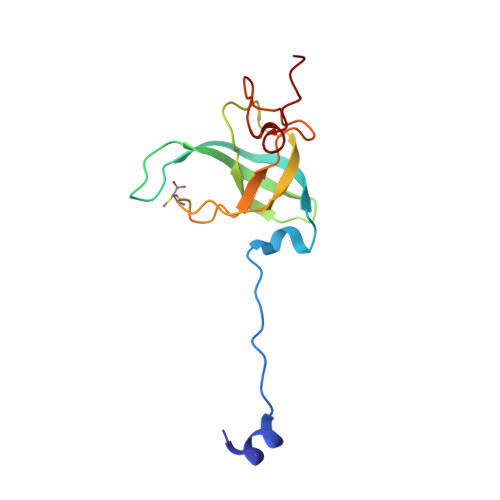
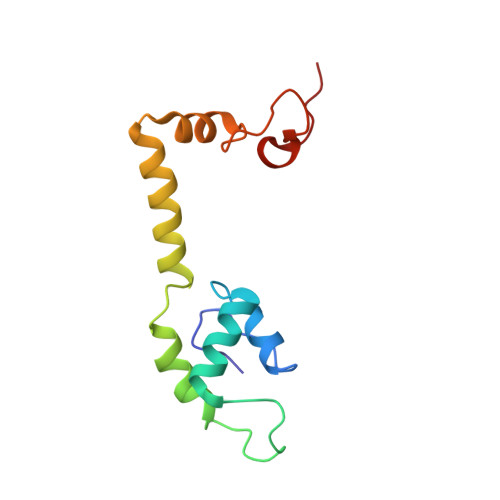
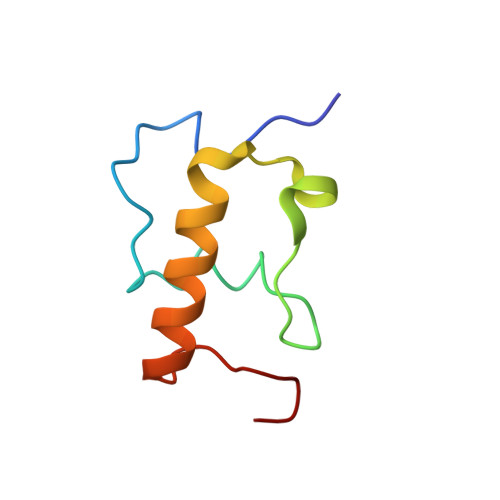
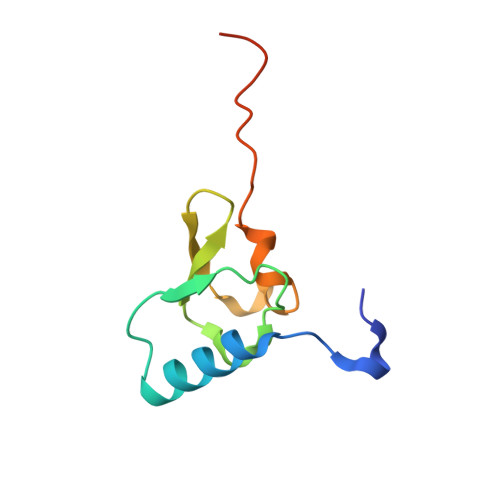
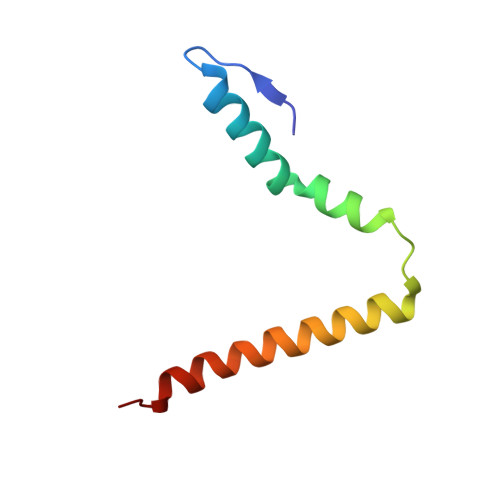
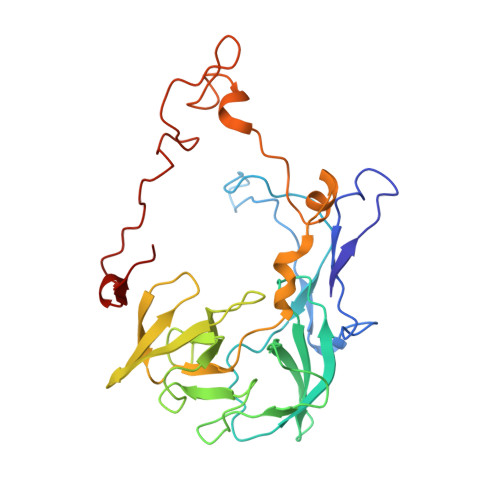
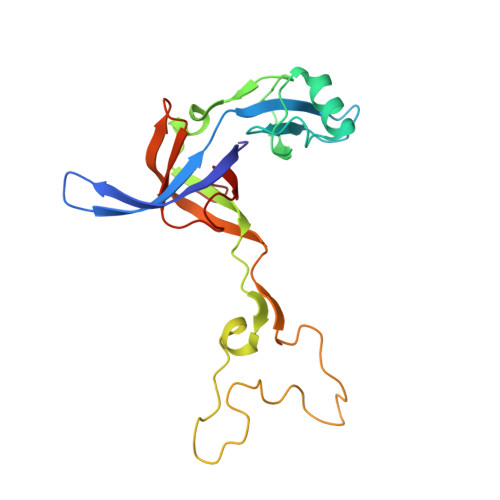
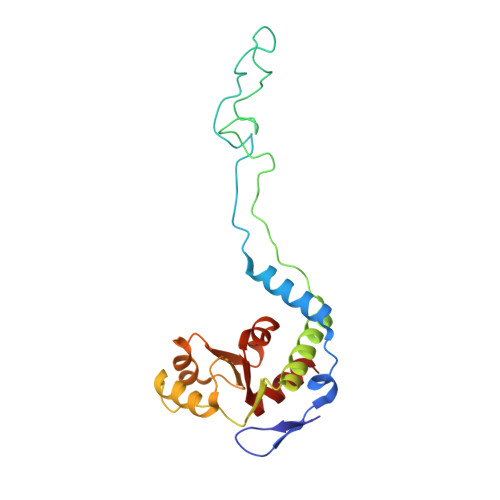
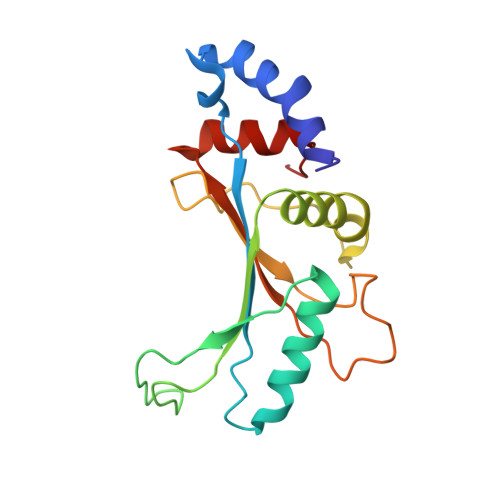
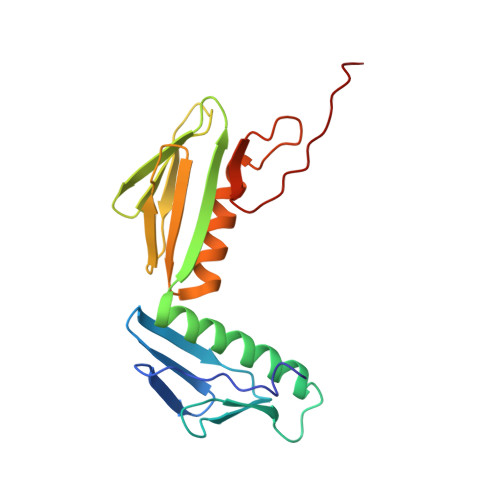
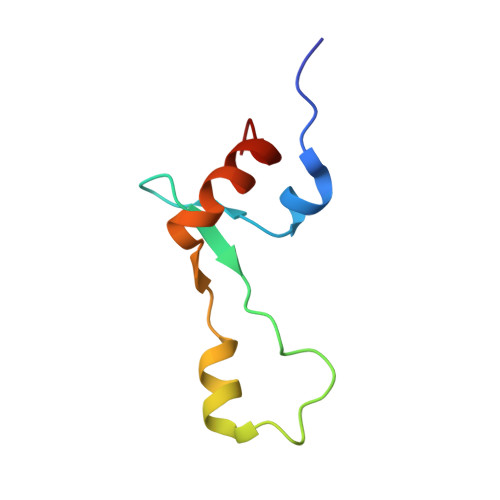
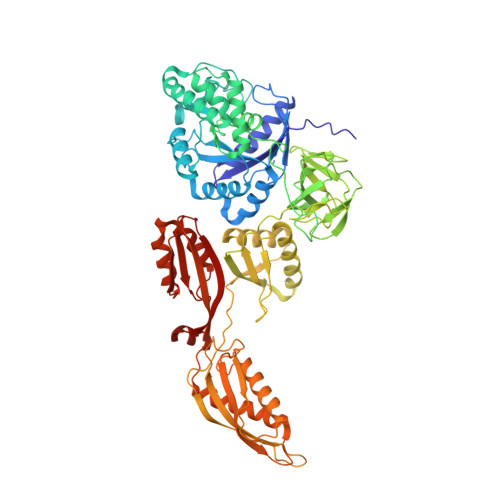
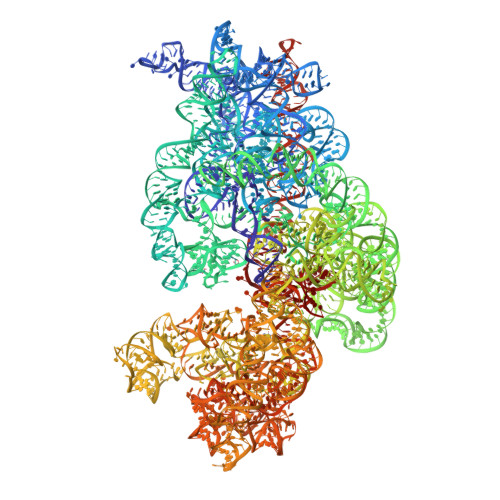
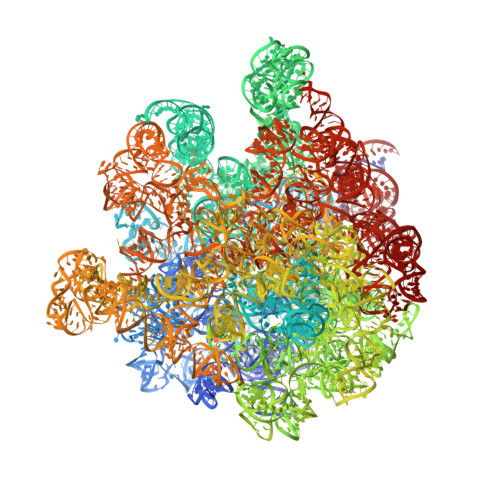
The cyclic octapeptide antibiotic argyrin B inhibits translation by trapping EF-G on the ribosome during translocation.
Wieland, M., Holm, M., Rundlet, E.J., Morici, M., Koller, T.O., Maviza, T.P., Pogorevc, D., Osterman, I.A., Muller, R., Blanchard, S.C., Wilson, D.N.(2022) Proc Natl Acad Sci U S A 119: e2114214119-e2114214119
- PubMed: 35500116 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2114214119
- Primary Citation Related Structures:
7OTC, 7UG7 - PubMed Abstract:
Argyrins are a family of naturally produced octapeptides that display promising antimicrobial activity against Pseudomonas aeruginosa. Argyrin B (ArgB) has been shown to interact with an elongated form of the translation elongation factor G (EF-G), leading to the suggestion that argyrins inhibit protein synthesis by interfering with EF-G binding to the ribosome. Here, using a combination of cryo-electron microscopy (cryo-EM) and single-molecule fluorescence resonance energy transfer (smFRET), we demonstrate that rather than interfering with ribosome binding, ArgB rapidly and specifically binds EF-G on the ribosome to inhibit intermediate steps of the translocation mechanism. Our data support that ArgB inhibits conformational changes within EF-G after GTP hydrolysis required for translocation and factor dissociation, analogous to the mechanism of fusidic acid, a chemically distinct antibiotic that binds a different region of EF-G. These findings shed light on the mechanism of action of the argyrin-class antibiotics on protein synthesis as well as the nature and importance of rate-limiting, intramolecular conformational events within the EF-G-bound ribosome during late-steps of translocation.
- Institute for Biochemistry and Molecular Biology, University of Hamburg, 20146 Hamburg, Germany.
Organizational Affiliation: