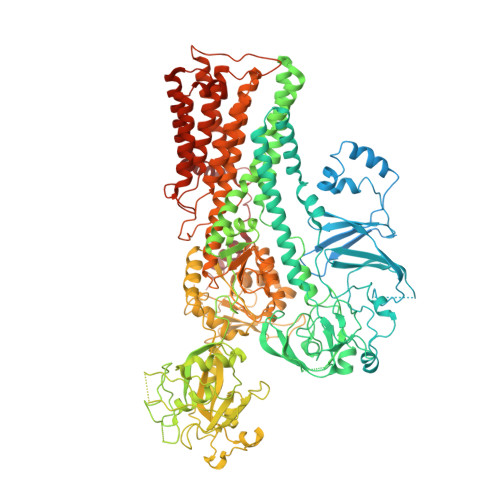
Structure and transport mechanism of P5B-ATPases.
Li, P., Wang, K., Salustros, N., Gronberg, C., Gourdon, P.(2021) Nat Commun 12: 3973-3973
- PubMed: 34172751 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-24148-y
- Primary Citation Related Structures:
7OP1, 7OP3, 7OP5, 7OP8 - PubMed Abstract:
In human cells, P5B-ATPases execute the active export of physiologically important polyamines such as spermine from lysosomes to the cytosol, a function linked to a palette of disorders. Yet, the overall shape of P5B-ATPases and the mechanisms of polyamine recognition, uptake and transport remain elusive. Here we describe a series of cryo-electron microscopy structures of a yeast homolog of human ATP13A2-5, Ypk9, determined at resolutions reaching 3.4 Å, and depicting three separate transport cycle intermediates, including spermine-bound conformations. Surprisingly, in the absence of cargo, Ypk9 rests in a phosphorylated conformation auto-inhibited by the N-terminus. Spermine uptake is accomplished through an electronegative cleft lined by transmembrane segments 2, 4 and 6. Despite the dramatically different nature of the transported cargo, these findings pinpoint shared principles of transport and regulation among the evolutionary related P4-, P5A- and P5B-ATPases. The data also provide a framework for analysis of associated maladies, such as Parkinson's disease.
- Department of Experimental Medical Science, Lund University, Lund, Sweden.
Organizational Affiliation: