Crystal Structure-Guided Design of Bisubstrate Inhibitors and Photoluminescent Probes for Protein Kinases of the PIM Family.
Nonga, O.E., Lavogina, D., Enkvist, E., Kestav, K., Chaikuad, A., Dixon-Clarke, S.E., Bullock, A.N., Kopanchuk, S., Ivan, T., Ekambaram, R., Viht, K., Knapp, S., Uri, A.(2021) Molecules 26
- PubMed: 34299628 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/molecules26144353
- Primary Citation Related Structures:
7OOV, 7OOW, 7OOX - PubMed Abstract:
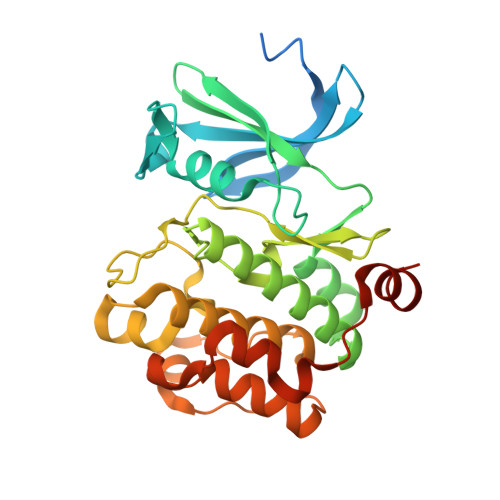
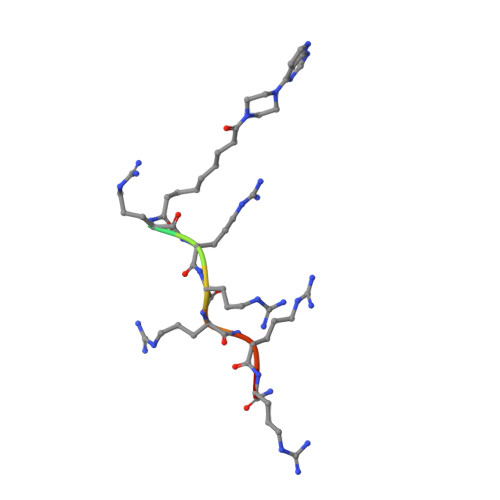
We performed an X-ray crystallographic study of complexes of protein kinase PIM-1 with three inhibitors comprising an adenosine mimetic moiety, a linker, and a peptide-mimetic (d-Arg) 6 fragment. Guided by the structural models, simplified chemical structures with a reduced number of polar groups and chiral centers were designed. The developed inhibitors retained low-nanomolar potency and possessed remarkable selectivity toward the PIM kinases. The new inhibitors were derivatized with biotin or fluorescent dye Cy5 and then applied for the detection of PIM kinases in biochemical solutions and in complex biological samples. The sandwich assay utilizing a PIM-2-selective detection antibody featured a low limit of quantification (44 pg of active recombinant PIM-2). Fluorescent probes were efficiently taken up by U2OS cells and showed a high extent of co-localization with PIM-1 fused with a fluorescent protein. Overall, the developed inhibitors and derivatives represent versatile chemical tools for studying PIM function in cellular systems in normal and disease physiology.
- Institute of Chemistry, University of Tartu, 14A Ravila St., 50411 Tartu, Estonia.
Organizational Affiliation: