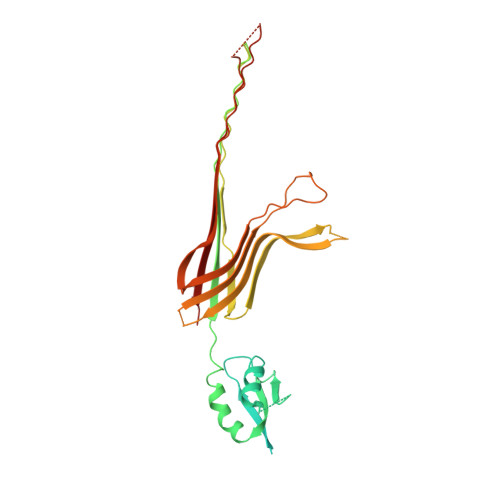
CryoEM structure of the outer membrane secretin channel pIV from the f1 filamentous bacteriophage.
Conners, R., McLaren, M., Lapinska, U., Sanders, K., Stone, M.R.L., Blaskovich, M.A.T., Pagliara, S., Daum, B., Rakonjac, J., Gold, V.A.M.(2021) Nat Commun 12: 6316-6316
- PubMed: 34728631 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-26610-3
- Primary Citation Related Structures:
7OFH - PubMed Abstract:
The Ff family of filamentous bacteriophages infect gram-negative bacteria, but do not cause lysis of their host cell. Instead, new virions are extruded via the phage-encoded pIV protein, which has homology with bacterial secretins. Here, we determine the structure of pIV from the f1 filamentous bacteriophage at 2.7 Å resolution by cryo-electron microscopy, the first near-atomic structure of a phage secretin. Fifteen f1 pIV subunits assemble to form a gated channel in the bacterial outer membrane, with associated soluble domains projecting into the periplasm. We model channel opening and propose a mechanism for phage egress. By single-cell microfluidics experiments, we demonstrate the potential for secretins such as pIV to be used as adjuvants to increase the uptake and efficacy of antibiotics in bacteria. Finally, we compare the f1 pIV structure to its homologues to reveal similarities and differences between phage and bacterial secretins.
- Living Systems Institute, University of Exeter, Exeter, UK.
Organizational Affiliation: