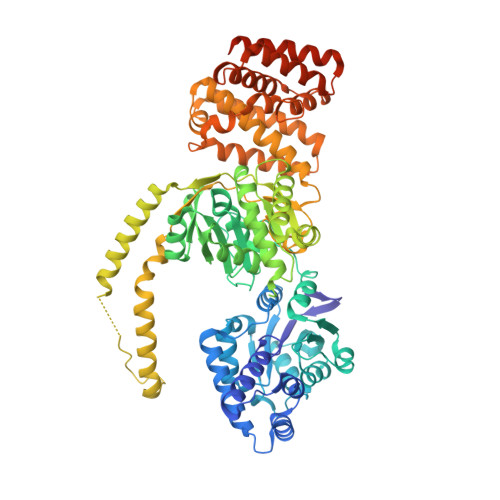
Unidirectional mannitol synthesis of Acinetobacter baumannii MtlD is facilitated by the helix-loop-helix-mediated dimer formation.
Tam, H.K., Konig, P., Himpich, S., Ngu, N.D., Abele, R., Muller, V., Pos, K.M.(2022) Proc Natl Acad Sci U S A 119: e2107994119-e2107994119
- PubMed: 35363566 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2107994119
- Primary Citation Related Structures:
7OCN, 7OCP, 7OCQ, 7OCR, 7OCS, 7OCT, 7OCU - PubMed Abstract:
Persistence of Acinetobacter baumannii in environments with low water activity is largely attributed to the biosynthesis of compatible solutes. Mannitol is one of the key compatible solutes in A. baumannii, and it is synthesized by a bifunctional mannitol-1-phosphate dehydrogenase/phosphatase (AbMtlD). AbMtlD catalyzes the conversion of fructose-6-phosphate to mannitol in two consecutive steps. Here, we report the crystal structure of dimeric AbMtlD, constituting two protomers each with a dehydrogenase and phosphatase domain. A proper assembly of AbMtlD dimer is facilitated by an intersection comprising a unique helix–loop–helix (HLH) domain. Reduction and dephosphorylation catalysis of fructose-6-phosphate to mannitol is dependent on the transient dimerization of AbMtlD. AbMtlD presents as a monomer under lower ionic strength conditions and was found to be mainly dimeric under high-salt conditions. The AbMtlD catalytic efficiency was markedly increased by cross-linking the protomers at the intersected HLH domain via engineered disulfide bonds. Inactivation of the AbMtlD phosphatase domain results in an intracellular accumulation of mannitol-1-phosphate in A. baumannii, leading to bacterial growth impairment upon salt stress. Taken together, our findings demonstrate that salt-induced dimerization of the bifunctional AbMtlD increases catalytic dehydrogenase and phosphatase efficiency, resulting in unidirectional catalysis of mannitol production.
- Institute of Biochemistry, Goethe University Frankfurt, Frankfurt am Main D-60438, Germany.
Organizational Affiliation: