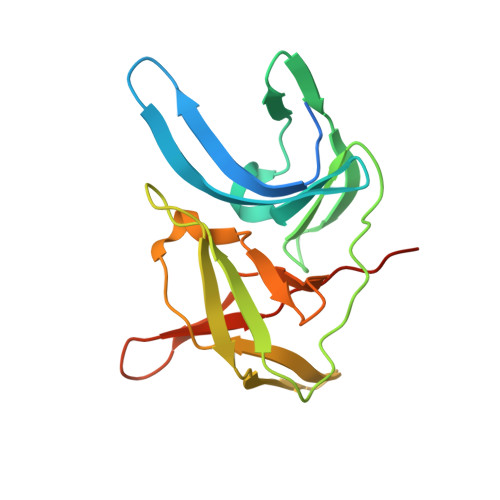
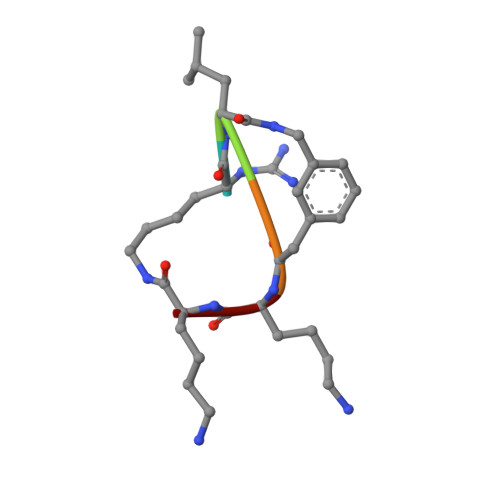
Structure-Based Optimization and Characterization of Macrocyclic Zika Virus NS2B-NS3 Protease Inhibitors.
Huber, S., Braun, N.J., Schmacke, L.C., Quek, J.P., Murra, R., Bender, D., Hildt, E., Luo, D., Heine, A., Steinmetzer, T.(2022) J Med Chem 65: 6555-6572
- PubMed: 35475620 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.1c01860
- Primary Citation Related Structures:
7O2M, 7O55, 7OBV, 7OC2, 7PFQ, 7PFY, 7PFZ, 7PG1, 7PGC, 7VLG, 7VLH, 7VLI - PubMed Abstract:
Zika virus (ZIKV) is a human pathogenic arbovirus. So far, neither a specific treatment nor a vaccination against ZIKV infections has been approved. Starting from our previously described lead structure, a series of 29 new macrocyclic inhibitors of the Zika virus protease containing different linker motifs have been synthesized. By selecting hydrophobic d-amino acids as part of the linker, numerous inhibitors with K i values < 5 nM were obtained. For 12 inhibitors, crystal structures in complex with the ZIKV protease up to 1.30 Å resolution were determined, which contribute to the understanding of the observed structure-activity relationship (SAR). In immunofluorescence assays, an antiviral effect was observed for compound 26 containing a d-homocyclohexylalanine residue in its linker segment. Due to its excellent selectivity profile and low cytotoxicity, this inhibitor scaffold could be a suitable starting point for the development of peptidic drugs against the Zika virus and related flaviviruses.
- Institute of Pharmaceutical Chemistry, Philipps University of Marburg, Marbacher Weg 6, 35032 Marburg, Germany.
Organizational Affiliation: