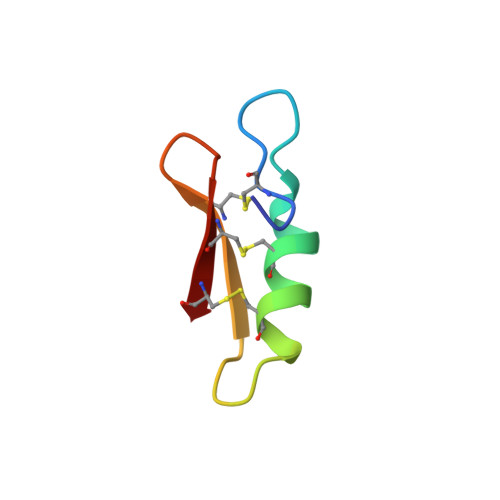
pH- and concentration-dependent supramolecular assembly of a fungal defensin plectasin variant into helical non-amyloid fibrils.
Pohl, C., Effantin, G., Kandiah, E., Meier, S., Zeng, G., Streicher, W., Segura, D.R., Mygind, P.H., Sandvang, D., Nielsen, L.A., Peters, G.H.J., Schoehn, G., Mueller-Dieckmann, C., Noergaard, A., Harris, P.(2022) Nat Commun 13: 3162-3162
- PubMed: 35672293 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-30462-w
- Primary Citation Related Structures:
7O76, 7OAE, 7OAG - PubMed Abstract:
Self-assembly and fibril formation play important roles in protein behaviour. Amyloid fibril formation is well-studied due to its role in neurodegenerative diseases and characterized by refolding of the protein into predominantly β-sheet form. However, much less is known about the assembly of proteins into other types of supramolecular structures. Using cryo-electron microscopy at a resolution of 1.97 Å, we show that a triple-mutant of the anti-microbial peptide plectasin, PPI42, assembles into helical non-amyloid fibrils. The in vitro anti-microbial activity was determined and shown to be enhanced compared to the wildtype. Plectasin contains a cysteine-stabilised α-helix-β-sheet structure, which remains intact upon fibril formation. Two protofilaments form a right-handed protein fibril. The fibril formation is reversible and follows sigmoidal kinetics with a pH- and concentration dependent equilibrium between soluble monomer and protein fibril. This high-resolution structure reveals that α/β proteins can natively assemble into fibrils.
- Novozymes A/S, Bagsvaerd, Denmark. christin.pohl@biochemistry.lu.se.
Organizational Affiliation: