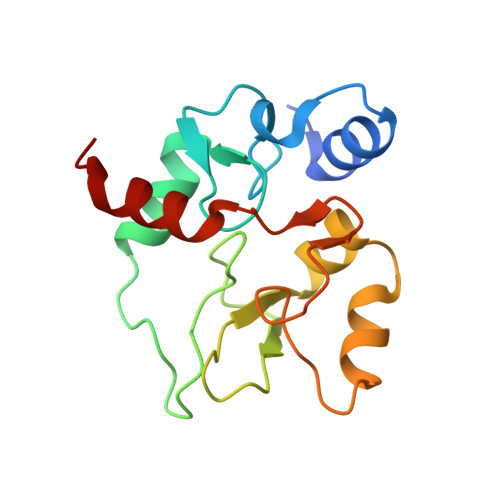
Structure of the DNMT3B ADD domain suggests the absence of a DNMT3A-like autoinhibitory mechanism.
Boyko, K., Arkova, O., Nikolaeva, A., Popov, V.O., Georgiev, P., Bonchuk, A.(2022) Biochem Biophys Res Commun 619: 124-129
- PubMed: 35760008 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2022.06.036
- Primary Citation Related Structures:
7O45 - PubMed Abstract:
De novo DNA methylation in early mammalian development depends on the activity of the DNMT3 methyltransferase family. An autoinhibitory mechanism involving the interaction between ADD and the catalytic domains of DNMT3A has been described. ADD is a zinc-coordinating histone-binding domain. The ADD domain of DNMT3A, when bound to a K4-unmethylated histone H3 tail, switches the enzyme to its catalytically active state. DNMT3B is another de novo methyltransferase enzyme with a more strict tissue- and stage-specific expression profile and a slightly different site specificity, lacking cooperative DNA methylation activity. Here, we obtained the crystal structure of the DNMT3B ADD domain, which demonstrated the extended conformation of the autoinhibitory loop even in the absence of the histone H3 tail. The lack of interaction between DNMT3B ADD and the methyltransferase domain was confirmed using an in vitro pull-down assay. The structural rearrangements in the loop also created an additional protein interaction interface leading to the formation of trimers in crystal, which may reflect their possible involvement in some unknown protein-protein interactions. Our results suggest that DNMT3B, in contrast to DNMT3A, has different modes of regulation of its activity that are independent of H3K4 methylation status.
- Department of the Control of Genetic Processes, Institute of Gene Biology Russian Academy of Sciences, 34/5 Vavilov St., Moscow, 119334, Russia; Bach Institute of Biochemistry, Research Center of Biotechnology Russian Academy of Sciences, Leninsky pr-t, 33, Bld. 2, Moscow, 119071, Russia.
Organizational Affiliation: