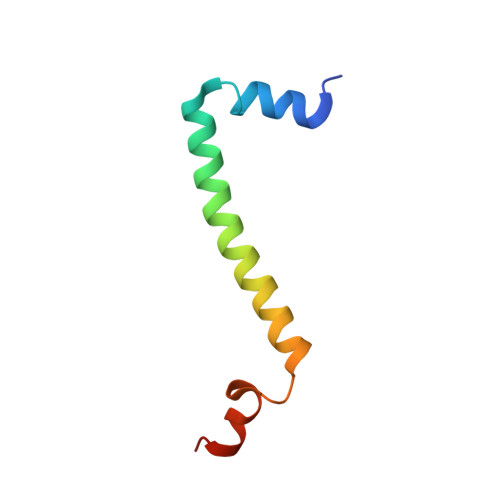
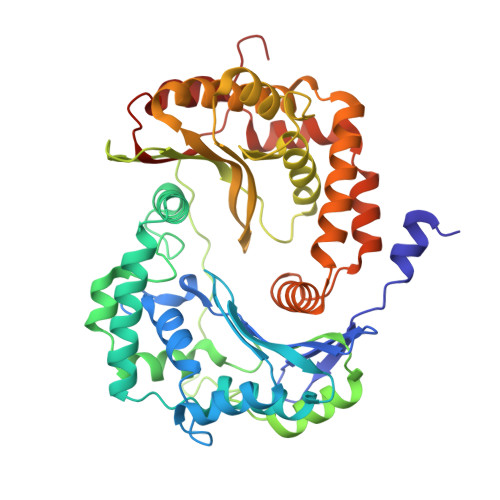
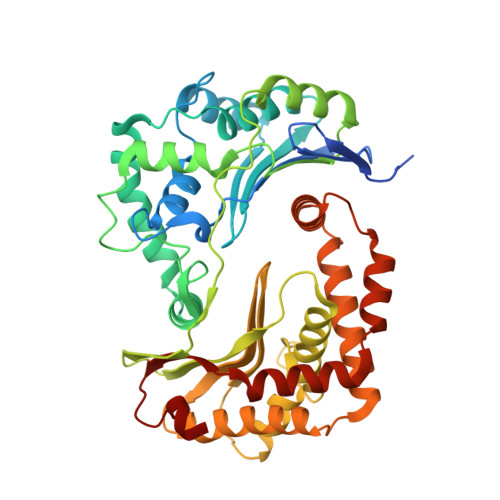
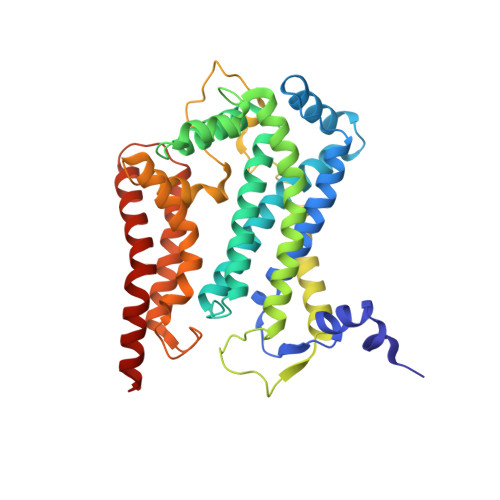
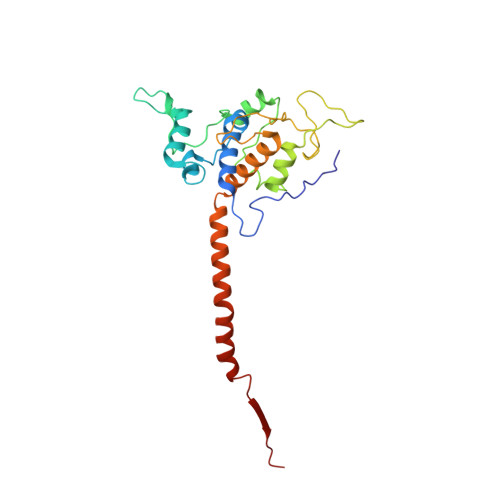
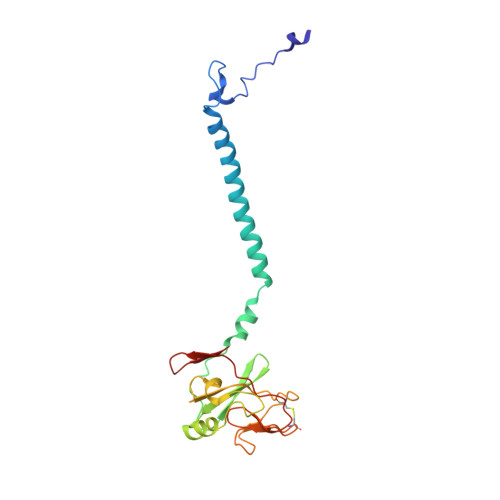
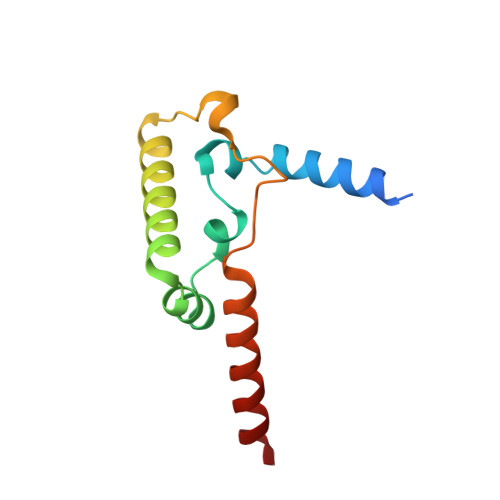
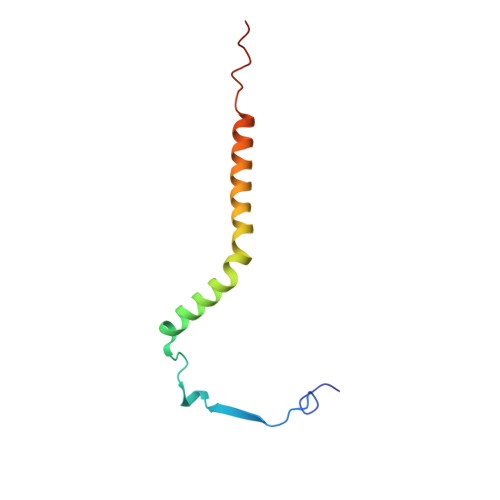
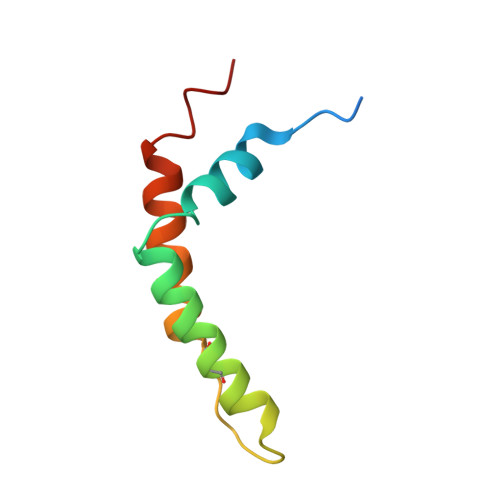
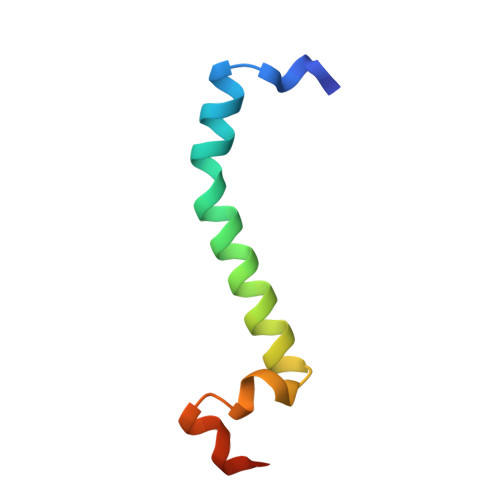
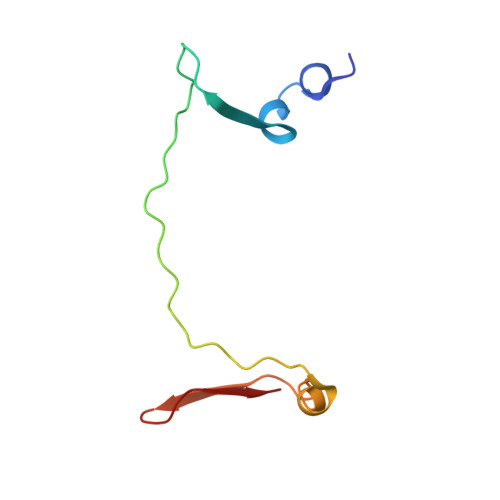
Structure and assembly of the mammalian mitochondrial supercomplex CIII 2 CIV.
Vercellino, I., Sazanov, L.A.(2021) Nature 598: 364-367
- PubMed: 34616041 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-021-03927-z
- Primary Citation Related Structures:
7O37, 7O3C, 7O3E, 7O3H - PubMed Abstract:
The enzymes of the mitochondrial electron transport chain are key players of cell metabolism. Despite being active when isolated, in vivo they associate into supercomplexes 1 , whose precise role is debated. Supercomplexes CIII 2 CIV 1-2 (refs. 2,3 ), CICIII 2 (ref. 4 ) and CICIII 2 CIV (respirasome) 5-10 exist in mammals, but in contrast to CICIII 2 and the respirasome, to date the only known eukaryotic structures of CIII 2 CIV 1-2 come from Saccharomyces cerevisiae 11,12 and plants 13 , which have different organization. Here we present the first, to our knowledge, structures of mammalian (mouse and ovine) CIII 2 CIV and its assembly intermediates, in different conformations. We describe the assembly of CIII 2 CIV from the CIII 2 precursor to the final CIII 2 CIV conformation, driven by the insertion of the N terminus of the assembly factor SCAF1 (ref. 14 ) deep into CIII 2 , while its C terminus is integrated into CIV. Our structures (which include CICIII 2 and the respirasome) also confirm that SCAF1 is exclusively required for the assembly of CIII 2 CIV and has no role in the assembly of the respirasome. We show that CIII 2 is asymmetric due to the presence of only one copy of subunit 9, which straddles both monomers and prevents the attachment of a second copy of SCAF1 to CIII 2 , explaining the presence of one copy of CIV in CIII 2 CIV in mammals. Finally, we show that CIII 2 and CIV gain catalytic advantage when assembled into the supercomplex and propose a role for CIII 2 CIV in fine tuning the efficiency of electron transfer in the electron transport chain.
- IV & LAS: IST Austria, Klosterneuburg, Austria.
Organizational Affiliation: