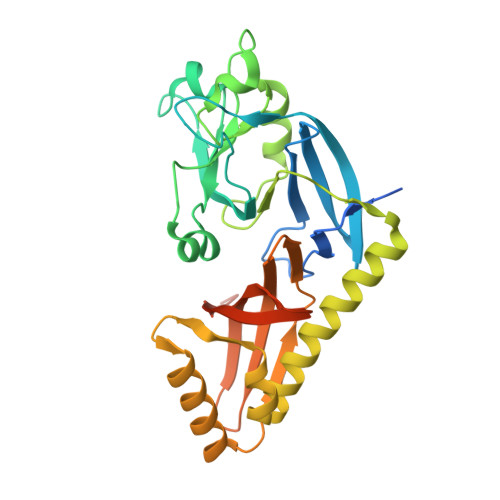
Asymmetric peptidoglycan editing generates cell curvature in Bdellovibrio predatory bacteria.
Banks, E.J., Valdivia-Delgado, M., Biboy, J., Wilson, A., Cadby, I.T., Vollmer, W., Lambert, C., Lovering, A.L., Sockett, R.E.(2022) Nat Commun 13: 1509-1509
- PubMed: 35314810 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-29007-y
- Primary Citation Related Structures:
7O21 - PubMed Abstract:
Peptidoglycan hydrolases contribute to the generation of helical cell shape in Campylobacter and Helicobacter bacteria, while cytoskeletal or periskeletal proteins determine the curved, vibrioid cell shape of Caulobacter and Vibrio. Here, we identify a peptidoglycan hydrolase in the vibrioid-shaped predatory bacterium Bdellovibrio bacteriovorus which invades and replicates within the periplasm of Gram-negative prey bacteria. The protein, Bd1075, generates cell curvature in B. bacteriovorus by exerting LD-carboxypeptidase activity upon the predator cell wall as it grows inside spherical prey. Bd1075 localizes to the outer convex face of B. bacteriovorus; this asymmetric localization requires a nuclear transport factor 2-like (NTF2) domain at the protein C-terminus. We solve the crystal structure of Bd1075, which is monomeric with key differences to other LD-carboxypeptidases. Rod-shaped Δbd1075 mutants invade prey more slowly than curved wild-type predators and stretch invaded prey from within. We therefore propose that the vibrioid shape of B. bacteriovorus contributes to predatory fitness.
- Medical School, School of Life Sciences, University of Nottingham, Queen's Medical Centre, Nottingham, NG7 2UH, UK.
Organizational Affiliation: