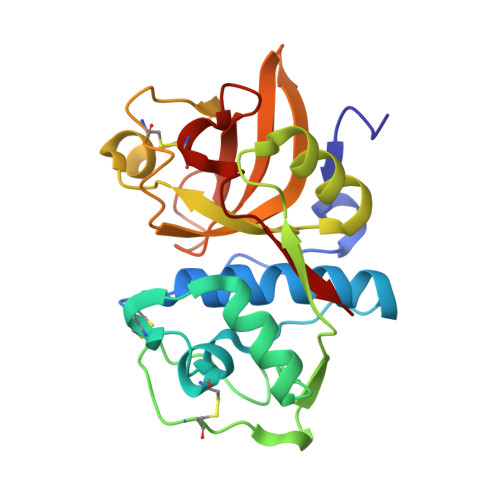
An Activity-Based Probe for Cathepsin K Imaging with Excellent Potency and Selectivity.
Lemke, C., Benysek, J., Brajtenbach, D., Breuer, C., Jilkova, A., Horn, M., Busa, M., Ulrychova, L., Illies, A., Kubatzky, K.F., Bartz, U., Mares, M., Gutschow, M.(2021) J Med Chem 64: 13793-13806
- PubMed: 34473502 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.1c01178
- Primary Citation Related Structures:
7NXL, 7NXM - PubMed Abstract:
The cysteine protease cathepsin K is a target for the treatment of diseases associated with high bone turnover. Cathepsin K is mainly expressed in osteoclasts and responsible for the destruction of the proteinaceous components of the bone matrix. We designed various fluorescent activity-based probes (ABPs) and their precursors that bind to and inactivate cathepsin K. ABP 25 exhibited extraordinary potency ( k inac / K i = 35,300 M -1 s -1 ) and selectivity for human cathepsin K. Crystal structures of cathepsin K in complex with ABP 25 and its nonfluorescent precursor 21 were determined to characterize the binding mode of this new type of acrylamide-based Michael acceptor with the particular orientation of the dibenzylamine moiety to the primed subsite region. The cyanine-5 containing probe 25 allowed for sensitive detection of cathepsin K, selective visualization in complex proteomes, and live cell imaging of a human osteosarcoma cell line, underlining its applicability in a pathophysiological environment.
- Pharmaceutical Institute, Pharmaceutical & Medicinal Chemistry, University of Bonn, An der Immenburg 4, Bonn 53121, Germany.
Organizational Affiliation: