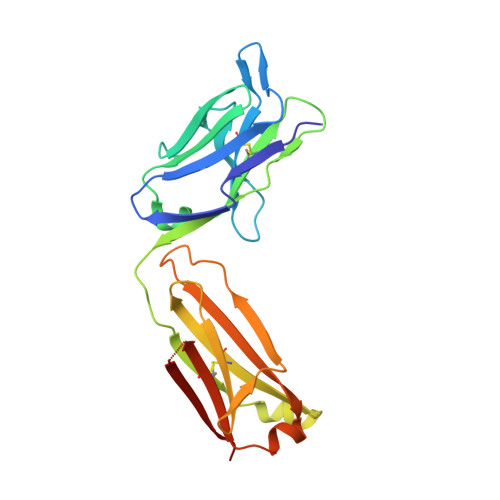
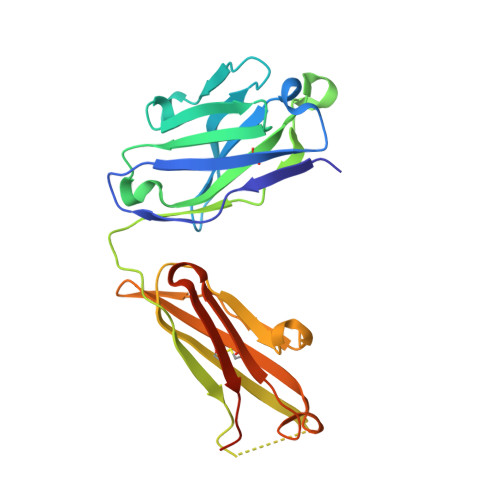
Structural basis of cytokine-mediated activation of ALK family receptors.
De Munck, S., Provost, M., Kurikawa, M., Omori, I., Mukohyama, J., Felix, J., Bloch, Y., Abdel-Wahab, O., Bazan, J.F., Yoshimi, A., Savvides, S.N.(2021) Nature 600: 143-147
- PubMed: 34646012 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-021-03959-5
- Primary Citation Related Structures:
7NWZ, 7NX0, 7NX1, 7NX2, 7NX3, 7NX4 - PubMed Abstract:
Anaplastic lymphoma kinase (ALK) 1 and the related leukocyte tyrosine kinase (LTK) 2 are recently deorphanized receptor tyrosine kinases 3 . Together with their activating cytokines, ALKAL1 and ALKAL2 4-6 (also called FAM150A and FAM150B or AUGβ and AUGα, respectively), they are involved in neural development 7 , cancer 7-9 and autoimmune diseases 10 . Furthermore, mammalian ALK recently emerged as a key regulator of energy expenditure and weight gain 11 , consistent with a metabolic role for Drosophila ALK 12 . Despite such functional pleiotropy and growing therapeutic relevance 13,14 , structural insights into ALK and LTK and their complexes with cognate cytokines have remained scarce. Here we show that the cytokine-binding segments of human ALK and LTK comprise a novel architectural chimera of a permuted TNF-like module that braces a glycine-rich subdomain featuring a hexagonal lattice of long polyglycine type II helices. The cognate cytokines ALKAL1 and ALKAL2 are monomeric three-helix bundles, yet their binding to ALK and LTK elicits similar dimeric assemblies with two-fold symmetry, that tent a single cytokine molecule proximal to the cell membrane. We show that the membrane-proximal EGF-like domain dictates the apparent cytokine preference of ALK. Assisted by these diverse structure-function findings, we propose a structural and mechanistic blueprint for complexes of ALK family receptors, and thereby extend the repertoire of ligand-mediated dimerization mechanisms adopted by receptor tyrosine kinases.
- Unit for Structural Biology, Department of Biochemistry and Microbiology, Ghent University, Ghent, Belgium.
Organizational Affiliation: