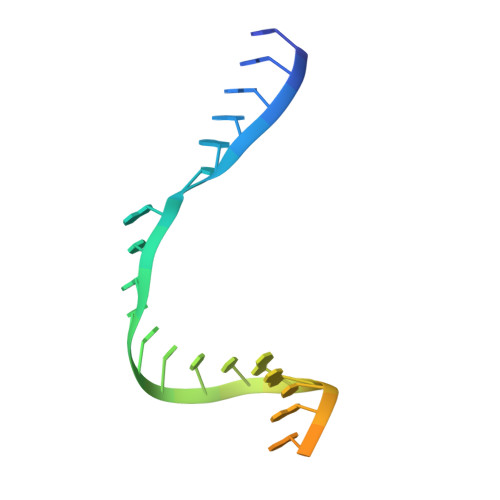
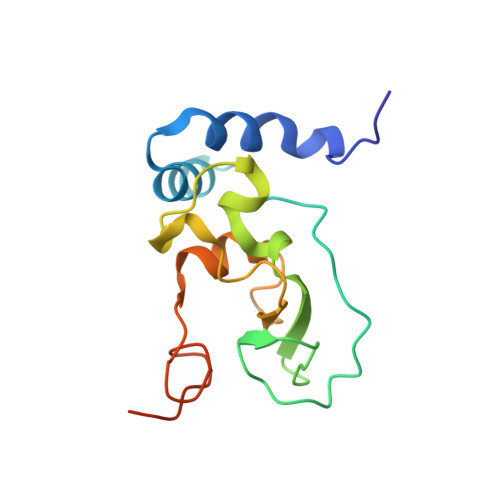
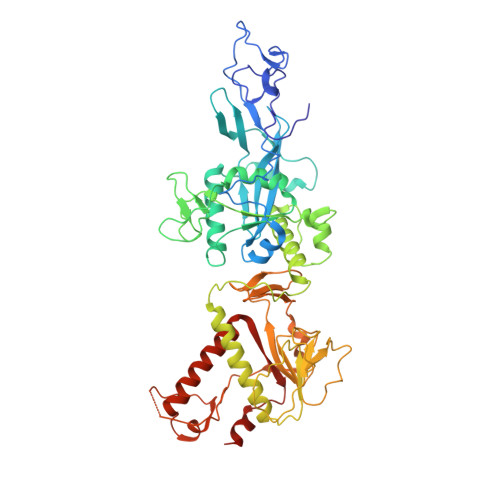
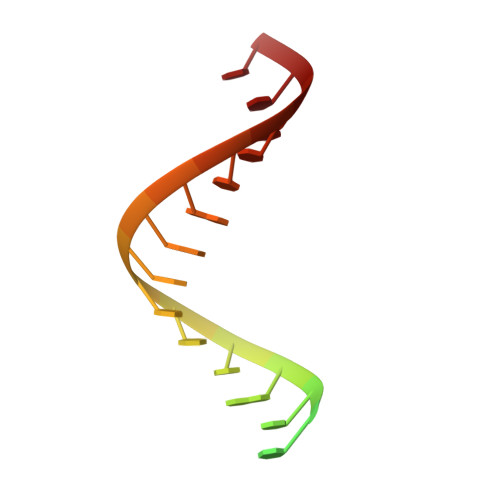
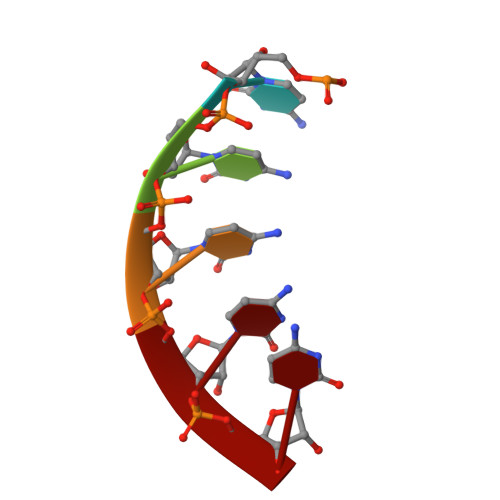
Structural basis of mismatch recognition by a SARS-CoV-2 proofreading enzyme.
Liu, C., Shi, W., Becker, S.T., Schatz, D.G., Liu, B., Yang, Y.(2021) Science 373: 1142-1146
- PubMed: 34315827 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.abi9310
- Primary Citation Related Structures:
7N0B, 7N0C, 7N0D - PubMed Abstract:
Coronavirus 3′-to-5′ exoribonuclease (ExoN), residing in the nonstructural protein (nsp) 10–nsp14 complex, boosts replication fidelity by proofreading RNA synthesis and is critical for the virus life cycle. ExoN also recognizes and excises nucleotide analog inhibitors incorporated into the nascent RNA, undermining the effectiveness of nucleotide analog–based antivirals. Here we present cryo–electron microscopy structures of both wild-type and mutant severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) nsp10-nsp14 in complex with an RNA substrate bearing a 3′-end mismatch at resolutions ranging from 2.5 to 3.9 angstroms. The structures reveal the molecular determinants of ExoN substrate specificity and offer insight into the molecular mechanisms of mismatch correction during coronavirus RNA synthesis. Our findings provide guidance for rational design of improved anticoronavirus therapies.
- Department of Immunobiology, Yale School of Medicine, New Haven, CT, USA.
Organizational Affiliation: