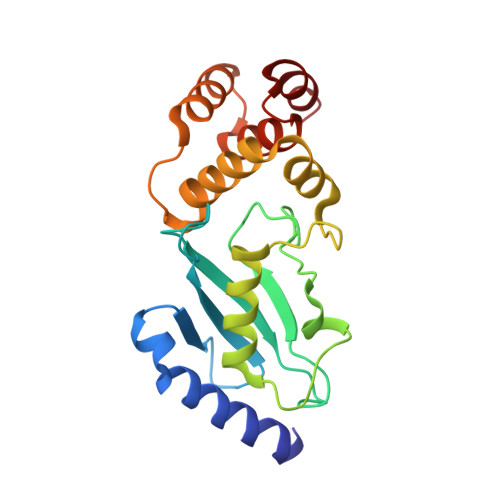
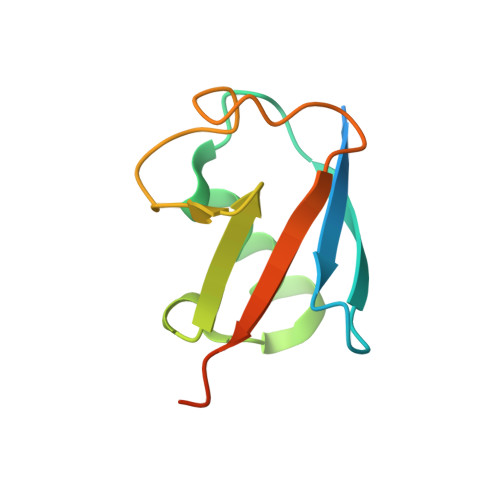
Identification of Ubiquitin Variants That Inhibit the E2 Ubiquitin Conjugating Enzyme, Ube2k.
Middleton, A.J., Teyra, J., Zhu, J., Sidhu, S.S., Day, C.L.(2021) ACS Chem Biol 16: 1745-1756
- PubMed: 34397214 Search on PubMed
- DOI: https://doi.org/10.1021/acschembio.1c00445
- Primary Citation Related Structures:
7MYF, 7MYH - PubMed Abstract:
Transfer of ubiquitin to substrate proteins regulates most processes in eukaryotic cells. E2 enzymes are a central component of the ubiquitin machinery, and generally determine the type of ubiquitin signal generated and thus the ultimate fate of substrate proteins. The E2, Ube2k, specifically builds degradative ubiquitin chains on diverse substrates. Here we have identified protein-based reagents, called ubiquitin variants (UbVs), that bind tightly and specifically to Ube2k. Crystal structures reveal that the UbVs bind to the E2 enzyme at a hydrophobic cleft that is distinct from the active site and previously identified ubiquitin binding sites. We demonstrate that the UbVs are potent inhibitors of Ube2k and block both ubiquitin charging of the E2 enzyme and E3-catalyzed ubiquitin transfer. The binding site of the UbVs suggests they directly clash with the ubiquitin activating enzyme, while potentially disrupting interactions with E3 ligases via allosteric effects. Our data reveal the first protein-based inhibitors of Ube2k and unveil a hydrophobic groove that could be an effective target for inhibiting Ube2k and other E2 enzymes.
- Department of Biochemistry, School of Biomedical Sciences, University of Otago, Dunedin 9054, New Zealand.
Organizational Affiliation: