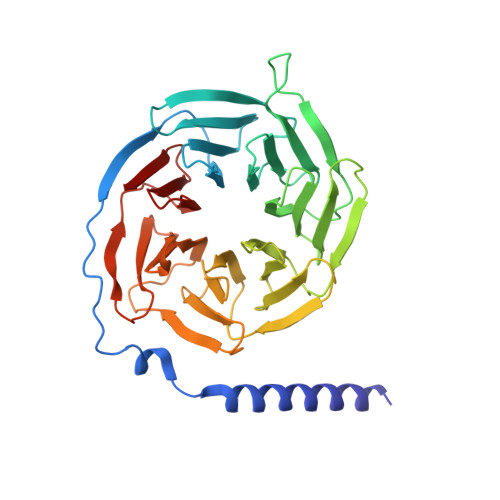
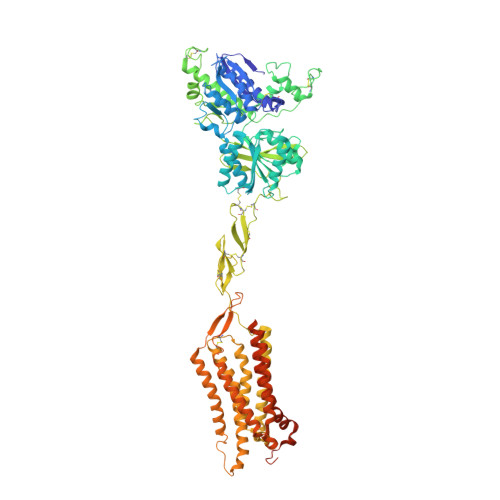
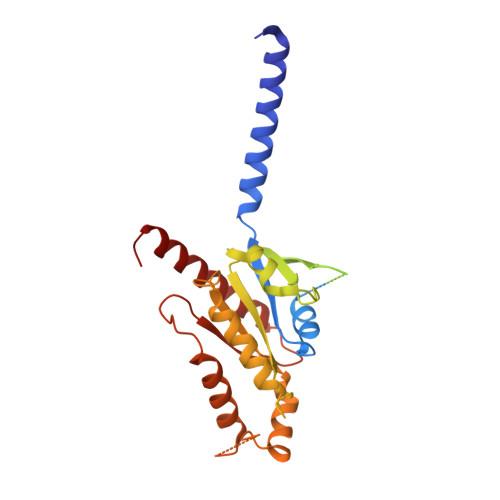
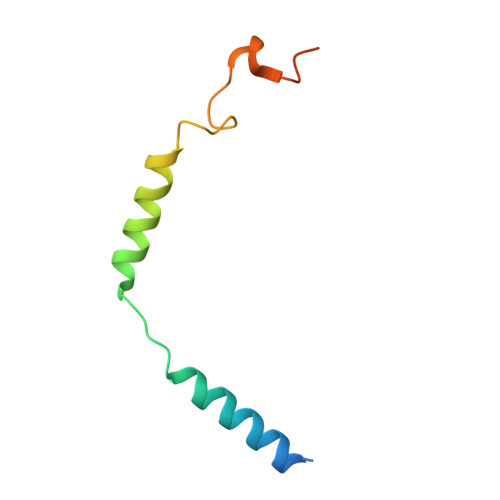
G-protein activation by a metabotropic glutamate receptor.
Seven, A.B., Barros-Alvarez, X., de Lapeyriere, M., Papasergi-Scott, M.M., Robertson, M.J., Zhang, C., Nwokonko, R.M., Gao, Y., Meyerowitz, J.G., Rocher, J.P., Schelshorn, D., Kobilka, B.K., Mathiesen, J.M., Skiniotis, G.(2021) Nature 595: 450-454
- PubMed: 34194039 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-021-03680-3
- Primary Citation Related Structures:
7MTQ, 7MTR, 7MTS - PubMed Abstract:
Family C G-protein-coupled receptors (GPCRs) operate as obligate dimers with extracellular domains that recognize small ligands, leading to G-protein activation on the transmembrane (TM) domains of these receptors by an unknown mechanism 1 . Here we show structures of homodimers of the family C metabotropic glutamate receptor 2 (mGlu2) in distinct functional states and in complex with heterotrimeric G i . Upon activation of the extracellular domain, the two transmembrane domains undergo extensive rearrangement in relative orientation to establish an asymmetric TM6-TM6 interface that promotes conformational changes in the cytoplasmic domain of one protomer. Nucleotide-bound G i can be observed pre-coupled to inactive mGlu2, but its transition to the nucleotide-free form seems to depend on establishing the active-state TM6-TM6 interface. In contrast to family A and B GPCRs, G-protein coupling does not involve the cytoplasmic opening of TM6 but is facilitated through the coordination of intracellular loops 2 and 3, as well as a critical contribution from the C terminus of the receptor. The findings highlight the synergy of global and local conformational transitions to facilitate a new mode of G-protein activation.
- Department of Molecular and Cellular Physiology, Stanford University School of Medicine, Stanford, CA, USA.
Organizational Affiliation: