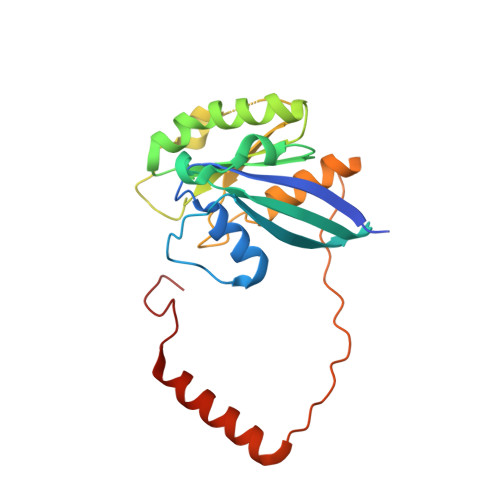
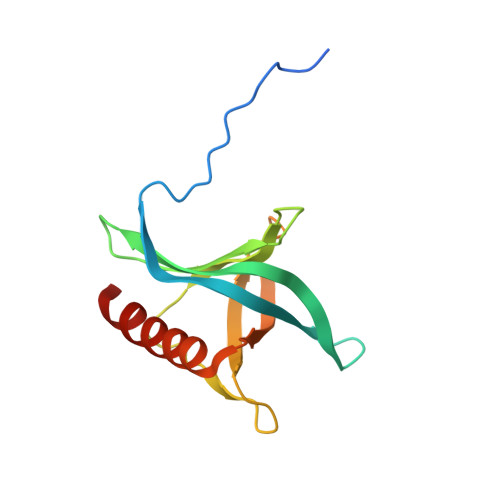
Architecture of the cytoplasmic face of the nuclear pore.
Bley, C.J., Nie, S., Mobbs, G.W., Petrovic, S., Gres, A.T., Liu, X., Mukherjee, S., Harvey, S., Huber, F.M., Lin, D.H., Brown, B., Tang, A.W., Rundlet, E.J., Correia, A.R., Chen, S., Regmi, S.G., Stevens, T.A., Jette, C.A., Dasso, M., Patke, A., Palazzo, A.F., Kossiakoff, A.A., Hoelz, A.(2022) Science 376: eabm9129-eabm9129
- PubMed: 35679405 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.abm9129
- Primary Citation Related Structures:
7MNI, 7MNJ, 7MNK, 7MNL, 7MNM, 7MNN, 7MNO, 7MNP, 7MNQ, 7MNR, 7MNS, 7MNT, 7MNU, 7MNV, 7MNW, 7MNX, 7MNY, 7MNZ, 7MO0, 7MO1, 7MO2, 7MO3, 7MO4, 7MO5, 7TBL, 7TBM - PubMed Abstract:
INTRODUCTION The subcellular compartmentalization of eukaryotic cells requires selective transport of folded proteins and protein-nucleic acid complexes. Embedded in nuclear envelope pores, which are generated by the circumscribed fusion of the inner and outer nuclear membranes, nuclear pore complexes (NPCs) are the sole bidirectional gateways for nucleocytoplasmic transport. The ~110-MDa human NPC is an ~1000-protein assembly that comprises multiple copies of ~34 different proteins, collectively termed nucleoporins. The symmetric core of the NPC is composed of an inner ring encircling the central transport channel and outer rings formed by Y‑shaped coat nucleoporin complexes (CNCs) anchored atop both sides of the nuclear envelope. The outer rings are decorated with compartment‑specific asymmetric nuclear basket and cytoplasmic filament nucleoporins, which establish transport directionality and provide docking sites for transport factors and the small guanosine triphosphatase Ran. The cytoplasmic filament nucleoporins also play an essential role in the irreversible remodeling of messenger ribonucleoprotein particles (mRNPs) as they exit the central transport channel. Unsurprisingly, the NPC's cytoplasmic face represents a hotspot for disease‑associated mutations and is commonly targeted by viral virulence factors. RATIONALE Previous studies established a near-atomic composite structure of the human NPC's symmetric core by combining (i) biochemical reconstitution to elucidate the interaction network between symmetric nucleoporins, (ii) crystal and single-particle cryo-electron microscopy structure determination of nucleoporins and nucleoporin complexes to reveal their three-dimensional shape and the molecular details of their interactions, (iii) quantitative docking in cryo-electron tomography (cryo-ET) maps of the intact human NPC to uncover nucleoporin stoichiometry and positioning, and (iv) cell‑based assays to validate the physiological relevance of the biochemical and structural findings. In this work, we extended our approach to the cytoplasmic filament nucleoporins to reveal the near-atomic architecture of the cytoplasmic face of the human NPC. RESULTS Using biochemical reconstitution, we elucidated the protein-protein and protein-RNA interaction networks of the human and Chaetomium thermophilum cytoplasmic filament nucleoporins, establishing an evolutionarily conserved heterohexameric cytoplasmic filament nucleoporin complex (CFNC) held together by a central heterotrimeric coiled‑coil hub that tethers two separate mRNP‑remodeling complexes. Further biochemical analysis and determination of a series of crystal structures revealed that the metazoan‑specific cytoplasmic filament nucleoporin NUP358 is composed of 16 distinct domains, including an N‑terminal S‑shaped α‑helical solenoid followed by a coiled‑coil oligomerization element, numerous Ran‑interacting domains, an E3 ligase domain, and a C‑terminal prolyl‑isomerase domain. Physiologically validated quantitative docking into cryo-ET maps of the intact human NPC revealed that pentameric NUP358 bundles, conjoined by the oligomerization element, are anchored through their N‑terminal domains to the central stalk regions of the CNC, projecting flexibly attached domains as far as ~600 Å into the cytoplasm. Using cell‑based assays, we demonstrated that NUP358 is dispensable for the architectural integrity of the assembled interphase NPC and RNA export but is required for efficient translation. After NUP358 assignment, the remaining 4-shaped cryo‑ET density matched the dimensions of the CFNC coiled‑coil hub, in close proximity to an outer-ring NUP93. Whereas the N-terminal NUP93 assembly sensor motif anchors the properly assembled related coiled‑coil channel nucleoporin heterotrimer to the inner ring, biochemical reconstitution confirmed that the NUP93 assembly sensor is reused in anchoring the CFNC to the cytoplasmic face of the human NPC. By contrast, two C. thermophilum CFNCs are anchored by a divergent mechanism that involves assembly sensors located in unstructured portions of two CNC nucleoporins. Whereas unassigned cryo‑ET density occupies the NUP358 and CFNC binding sites on the nuclear face, docking of the nuclear basket component ELYS established that the equivalent position on the cytoplasmic face is unoccupied, suggesting that mechanisms other than steric competition promote asymmetric distribution of nucleoporins. CONCLUSION We have substantially advanced the biochemical and structural characterization of the asymmetric nucleoporins' architecture and attachment at the cytoplasmic and nuclear faces of the NPC. Our near‑atomic composite structure of the human NPC's cytoplasmic face provides a biochemical and structural framework for elucidating the molecular basis of mRNP remodeling, viral virulence factor interference with NPC function, and the underlying mechanisms of nucleoporin diseases at the cytoplasmic face of the NPC. [Figure: see text].
- Division of Chemistry and Chemical Engineering, California Institute of Technology, 1200 East California Boulevard, Pasadena, CA 91125, USA.
Organizational Affiliation: