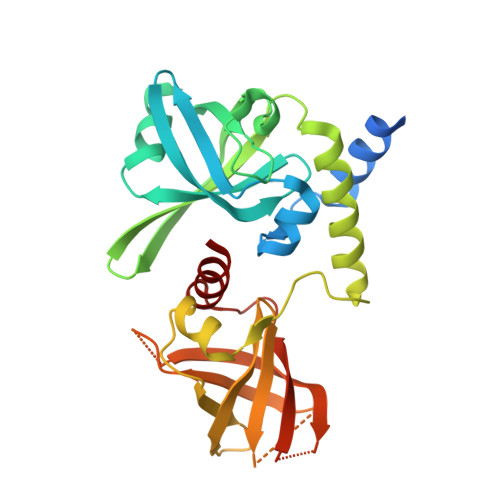
High-resolution crystal structure of the Borreliella burgdorferi PlzA protein in complex with c-di-GMP: new insights into the interaction of c-di-GMP with the novel xPilZ domain.
Singh, A., Izac, J.R., Schuler, E.J.A., Patel, D.T., Davies, C., Marconi, R.T.(2021) Pathog Dis 79
- PubMed: 34117751 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/femspd/ftab030
- Primary Citation Related Structures:
7MIE - PubMed Abstract:
In the tick-borne pathogens, Borreliella burgdorferi and Borrelia hermsii, c-di-GMP is produced by a single diguanylate cyclase (Rrp1). In these pathogens, the Plz proteins (PlzA, B and C) are the only c-di-GMP receptors identified to date and PlzA is the sole c-di-GMP receptor found in all Borreliella isolates. Bioinformatic analyses suggest that PlzA has a unique PilZN3-PilZ architecture with the relatively uncommon xPilZ domain. Here, we present the crystal structure of PlzA in complex with c-di-GMP (1.6 Å resolution). This is the first structure of a xPilz domain in complex with c-di-GMP to be determined. PlzA has a two-domain structure, where each domain comprises topologically equivalent PilZ domains with minimal sequence identity but remarkable structural similarity. The c-di-GMP binding site is formed by the linker connecting the two domains. While the structure of apo PlzA could not be determined, previous fluorescence resonance energy transfer data suggest that apo and holo forms of the protein are structurally distinct. The information obtained from this study will facilitate ongoing efforts to identify the molecular mechanisms of PlzA-mediated regulation in ticks and mammals.
- Department of Biochemistry and Molecular Biology, Medical University of South Carolina, Charleston, South Carolina 29425, USA.
Organizational Affiliation: