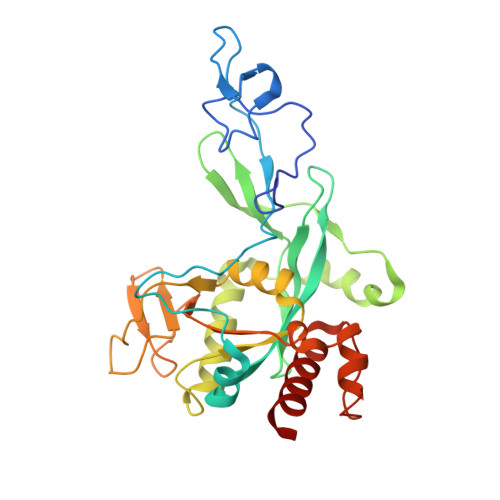
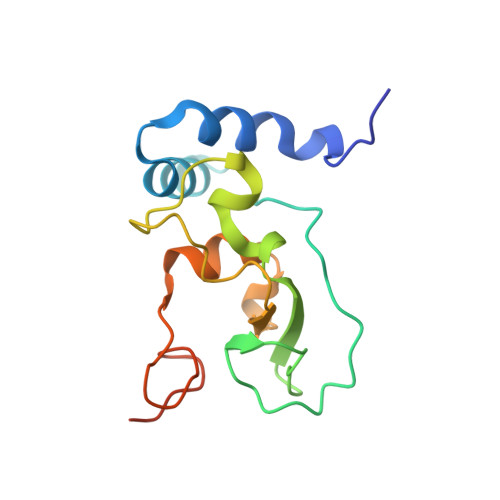
Structure and dynamics of SARS-CoV-2 proofreading exoribonuclease ExoN.
Moeller, N.H., Shi, K., Demir, O., Belica, C., Banerjee, S., Yin, L., Durfee, C., Amaro, R.E., Aihara, H.(2022) Proc Natl Acad Sci U S A 119
- PubMed: 35165203 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2106379119
- Primary Citation Related Structures:
7MC5, 7MC6 - PubMed Abstract:
High-fidelity replication of the large RNA genome of coronaviruses (CoVs) is mediated by a 3'-to-5' exoribonuclease (ExoN) in nonstructural protein 14 (nsp14), which excises nucleotides including antiviral drugs misincorporated by the low-fidelity viral RNA-dependent RNA polymerase (RdRp) and has also been implicated in viral RNA recombination and resistance to innate immunity. Here, we determined a 1.6-Å resolution crystal structure of severe acute respiratory syndrome CoV 2 (SARS-CoV-2) ExoN in complex with its essential cofactor, nsp10. The structure shows a highly basic and concave surface flanking the active site, comprising several Lys residues of nsp14 and the N-terminal amino group of nsp10. Modeling suggests that this basic patch binds to the template strand of double-stranded RNA substrates to position the 3' end of the nascent strand in the ExoN active site, which is corroborated by mutational and computational analyses. We also show that the ExoN activity can rescue a stalled RNA primer poisoned with sofosbuvir and allow RdRp to continue its extension in the presence of the chain-terminating drug, biochemically recapitulating proofreading in SARS-CoV-2 replication. Molecular dynamics simulations further show remarkable flexibility of multidomain nsp14 and suggest that nsp10 stabilizes ExoN for substrate RNA binding to support its exonuclease activity. Our high-resolution structure of the SARS-CoV-2 ExoN-nsp10 complex serves as a platform for future development of anticoronaviral drugs or strategies to attenuate the viral virulence.
- Department of Biochemistry, Molecular Biology and Biophysics, University of Minnesota, Minneapolis, MN 55455.
Organizational Affiliation: