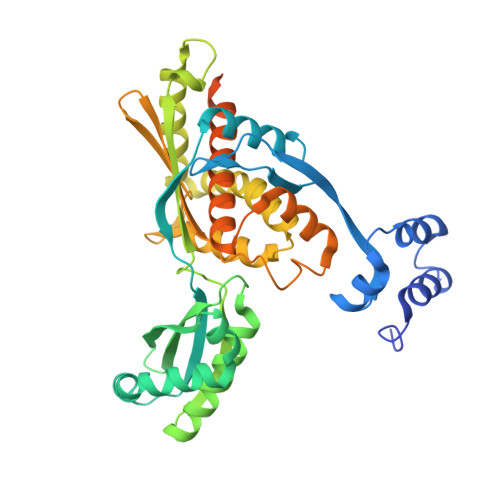
Targeting Enterococcus faecalis HMG-CoA reductase with a non-statin inhibitor.
Bose, S., Steussy, C.N., Lopez-Perez, D., Schmidt, T., Kulathunga, S.C., Seleem, M.N., Lipton, M., Mesecar, A.D., Rodwell, V.W., Stauffacher, C.V.(2023) Commun Biol 6: 360-360
- PubMed: 37012403 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-023-04639-y
- Primary Citation Related Structures:
7M66 - PubMed Abstract:
HMG-CoA reductase (HMGR), a rate-limiting enzyme of the mevalonate pathway in Gram-positive pathogenic bacteria, is an attractive target for development of novel antibiotics. In this study, we report the crystal structures of HMGR from Enterococcus faecalis (efHMGR) in the apo and liganded forms, highlighting several unique features of this enzyme. Statins, which inhibit the human enzyme with nanomolar affinity, perform poorly against the bacterial HMGR homologs. We also report a potent competitive inhibitor (Chembridge2 ID 7828315 or compound 315) of the efHMGR enzyme identified by a high-throughput, in-vitro screening. The X-ray crystal structure of efHMGR in complex with 315 was determined to 1.27 Å resolution revealing that the inhibitor occupies the mevalonate-binding site and interacts with several key active site residues conserved among bacterial homologs. Importantly, 315 does not inhibit the human HMGR. Our identification of a selective, non-statin inhibitor of bacterial HMG-CoA reductases will be instrumental in lead optimization and development of novel antibacterial drug candidates.
- Department of Biological Sciences, Purdue University, 915 West State Street, West Lafayette, IN, 47907, USA.
Organizational Affiliation: