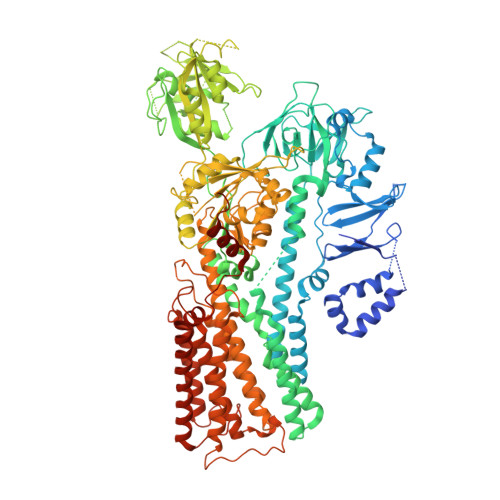
Structural mechanisms for gating and ion selectivity of the human polyamine transporter ATP13A2.
Tillinghast, J., Drury, S., Bowser, D., Benn, A., Lee, K.P.K.(2021) Mol Cell 81: 4650-4662.e4
- PubMed: 34715014 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2021.10.002
- Primary Citation Related Structures:
7M5V, 7M5X, 7M5Y - PubMed Abstract:
Mutations in ATP13A2, also known as PARK9, cause a rare monogenic form of juvenile-onset Parkinson's disease named Kufor-Rakeb syndrome and other neurodegenerative diseases. ATP13A2 encodes a neuroprotective P5B P-type ATPase highly enriched in the brain that mediates selective import of spermine ions from lysosomes into the cytosol via an unknown mechanism. Here we present three structures of human ATP13A2 bound to an ATP analog or to spermine in the presence of phosphomimetics determined by cryoelectron microscopy. ATP13A2 autophosphorylation opens a lysosome luminal gate to reveal a narrow lumen access channel that holds a spermine ion in its entrance. ATP13A2's architecture suggests physical principles underlying selective polyamine transport and anticipates a "pump-channel" intermediate that could function as a counter-cation conduit to facilitate lysosome acidification. Our findings establish a firm foundation to understand ATP13A2 mutations associated with disease and bring us closer to realizing ATP13A2's potential in neuroprotective therapy.
- Department of Cellular and Molecular Physiology, Pennsylvania State University College of Medicine, The Pennsylvania State University, Hershey, PA 17033, USA.
Organizational Affiliation: