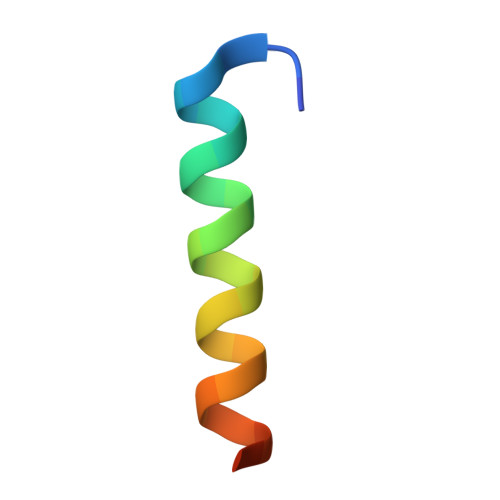
Structural basis of BAK activation in mitochondrial apoptosis initiation.
Singh, G., Guibao, C.D., Seetharaman, J., Aggarwal, A., Grace, C.R., McNamara, D.E., Vaithiyalingam, S., Waddell, M.B., Moldoveanu, T.(2022) Nat Commun 13: 250-250
- PubMed: 35017502 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-27851-y
- Primary Citation Related Structures:
7M5A, 7M5B, 7M5C - PubMed Abstract:
BCL-2 proteins regulate mitochondrial poration in apoptosis initiation. How the pore-forming BCL-2 Effector BAK is activated remains incompletely understood mechanistically. Here we investigate autoactivation and direct activation by BH3-only proteins, which cooperate to lower BAK threshold in membrane poration and apoptosis initiation. We define in trans BAK autoactivation as the asymmetric "BH3-in-groove" triggering of dormant BAK by active BAK. BAK autoactivation is mechanistically similar to direct activation. The structure of autoactivated BAK BH3-BAK complex reveals the conformational changes leading to helix α1 destabilization, which is a hallmark of BAK activation. Helix α1 is destabilized and restabilized in structures of BAK engaged by rationally designed, high-affinity activating and inactivating BID-like BH3 ligands, respectively. Altogether our data support the long-standing hit-and-run mechanism of BAK activation by transient binding of BH3-only proteins, demonstrating that BH3-induced structural changes are more important in BAK activation than BH3 ligand affinity.
- Department of Structural Biology, St. Jude Children's Research Hospital, Memphis, TN, USA.
Organizational Affiliation: