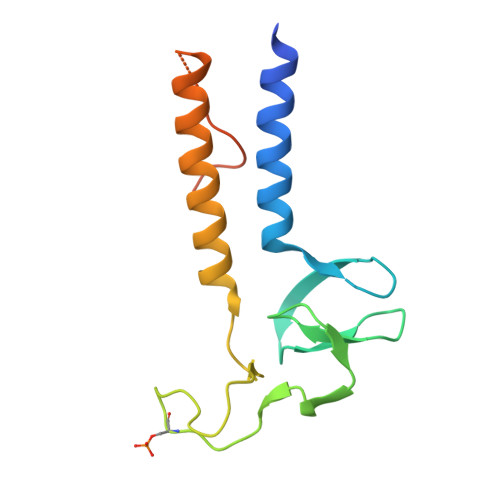
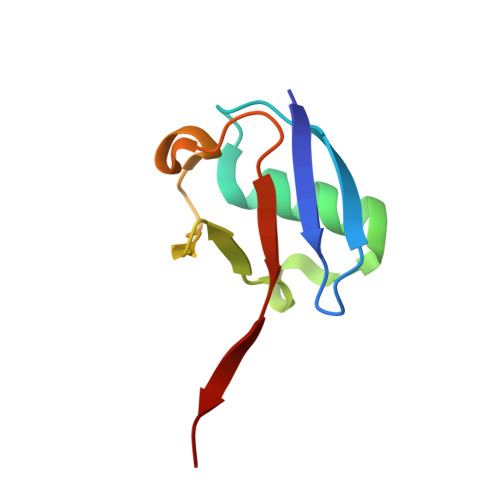
Structural basis of K63-ubiquitin chain formation by the Gordon-Holmes syndrome RBR E3 ubiquitin ligase RNF216.
Cotton, T.R., Cobbold, S.A., Bernardini, J.P., Richardson, L.W., Wang, X.S., Lechtenberg, B.C.(2022) Mol Cell 82: 598-615.e8
- PubMed: 34998453 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2021.12.005
- Primary Citation Related Structures:
7M4M, 7M4N, 7M4O - PubMed Abstract:
An increasing number of genetic diseases are linked to deregulation of E3 ubiquitin ligases. Loss-of-function mutations in the RING-between-RING (RBR) family E3 ligase RNF216 (TRIAD3) cause Gordon-Holmes syndrome (GHS) and related neurodegenerative diseases. Functionally, RNF216 assembles K63-linked ubiquitin chains and has been implicated in regulation of innate immunity signaling pathways and synaptic plasticity. Here, we report crystal structures of key RNF216 reaction states including RNF216 in complex with ubiquitin and its reaction product, K63 di-ubiquitin. Our data provide a molecular explanation for chain-type specificity and reveal the molecular basis for disruption of RNF216 function by pathogenic GHS mutations. Furthermore, we demonstrate how RNF216 activity and chain-type specificity are regulated by phosphorylation and that RNF216 is allosterically activated by K63-linked di-ubiquitin. These molecular insights expand our understanding of RNF216 function and its role in disease and further define the mechanistic diversity of the RBR E3 ligase family.
- Ubiquitin Signalling Division, The Walter and Eliza Hall Institute of Medical Research, 1G Royal Parade, Parkville, VIC 3052, Australia; Department of Medical Biology, The University of Melbourne, Parkville, VIC 3010, Australia.
Organizational Affiliation: