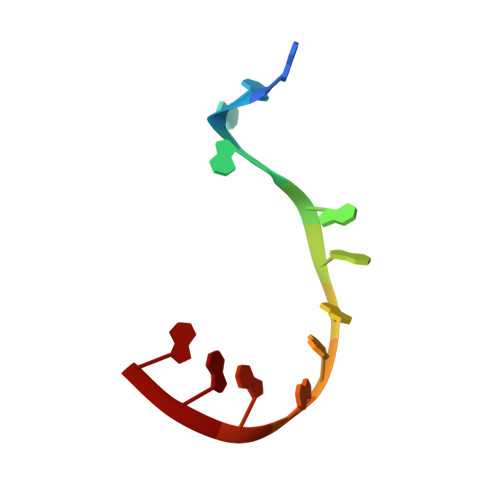
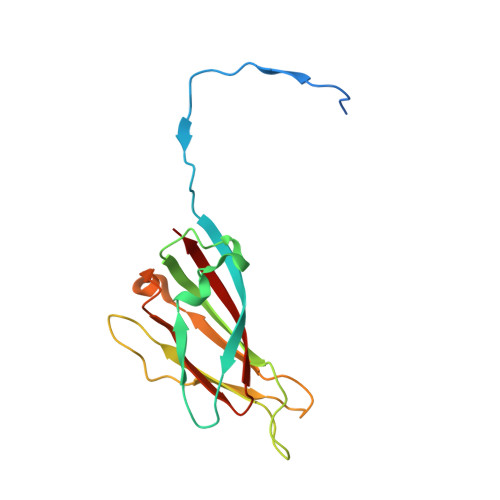
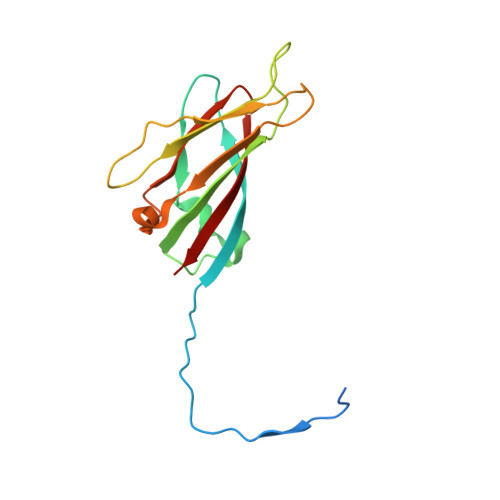
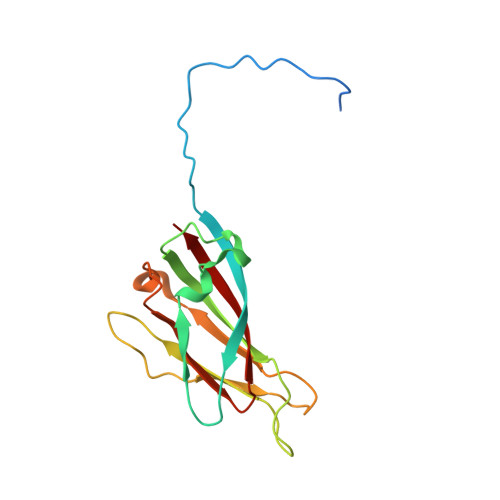
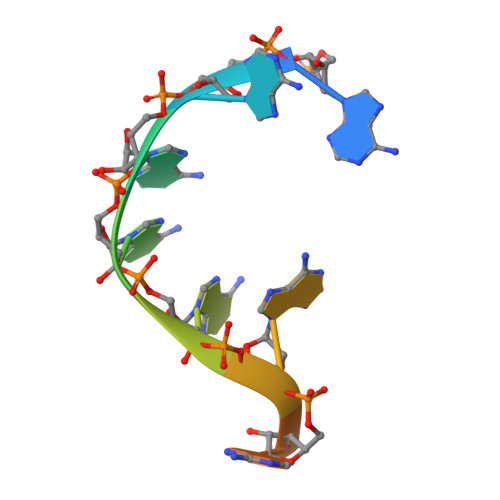
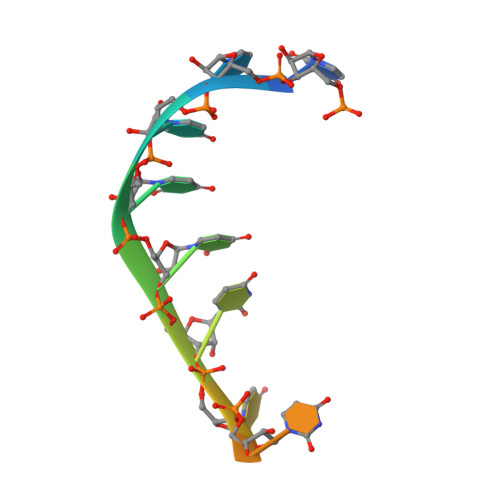
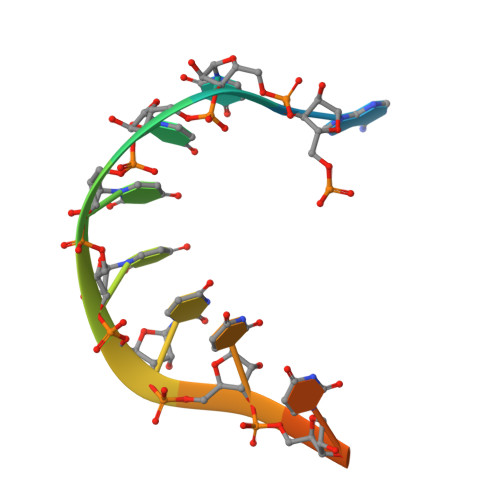
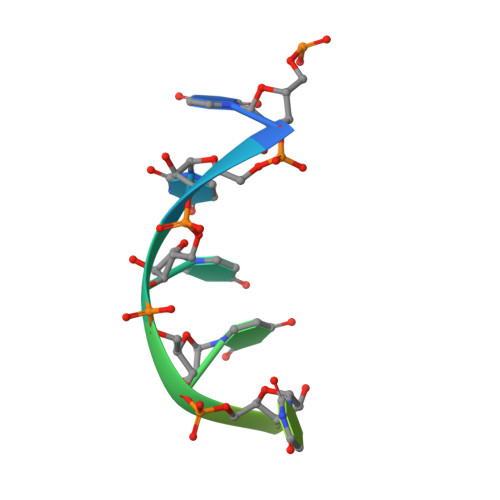
Structures of additional crystal forms of Satellite tobacco mosaic virus grown from a variety of salts.
McPherson, A.(2021) Acta Crystallogr F Struct Biol Commun 77: 473-483
- PubMed: 34866603 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X21011547
- Primary Citation Related Structures:
5BKL, 5BKN, 5BKQ, 7M2T, 7M2V, 7M3R, 7M3T, 7M50, 7M54, 7M57 - PubMed Abstract:
The structures of new crystal forms of Satellite tobacco mosaic virus (STMV) are described. These belong to space groups I2, P2 1 2 1 2 (a low-resolution form), R3 (H3) and P23. The R3 crystals are 50%/50% twinned, as are two instances of the P23 crystals. The I2 and P2 1 2 1 2 crystals were grown from ammonium sulfate solutions, as was one crystal in space group P23, while the R3 and the other P23 crystals were grown from sodium chloride, sodium bromide and sodium nitrate. The monoclinic and orthorhombic crystals have half a virus particle as the asymmetric unit, while the rhombohedral and cubic crystals have one third of a virus particle. RNA segments organized about the icosahedral twofold axes were present in crystals grown from ammonium sulfate and sodium chloride, as in the canonical I222 crystals (PDB entry 4oq8), but were not observed in crystals grown from sodium bromide and sodium nitrate. Bromide and nitrate ions generally replaced the RNA phosphates present in the I222 crystals, including the phosphates seen on fivefold axes, and were also found at threefold vertices in both the rhombohedral and cubic forms. An additional anion was also found on the fivefold axis 5 Å from the first anion, and slightly outside the capsid in crystals grown from sodium chloride, sodium bromide and sodium nitrate, suggesting that the path along the symmetry axis might be an ion channel. The electron densities for RNA strands at individual icosahedral dyads, as well as at the amino-terminal peptides of protein subunits, exhibited a diversity of orientations, in particular the residues at the ends.
- Department of Molecular Biology and Biochemistry, University of California, 530A Steinhaus Hall, Irvine, CA 92697-3900, USA.
Organizational Affiliation: