The crystal structure of Papain-Like Protease of SARS CoV-2, C111S mutant, in complex with ebselen
Osipiuk, J., Tesar, C., Endres, M., Maltseva, N., Joachimiak, A., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Papain-like protease | 318 | Severe acute respiratory syndrome coronavirus 2 | Mutation(s): 1 EC: 3.4.22 | 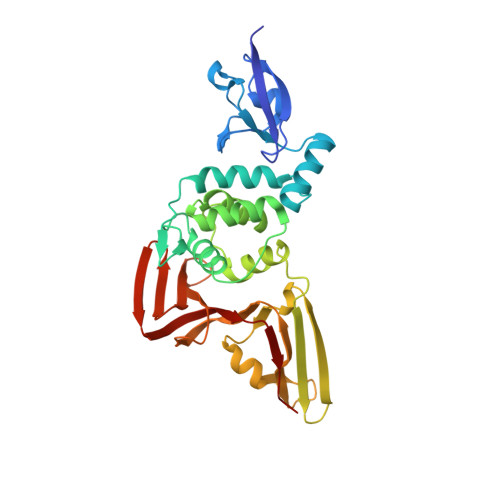 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0DTD1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 7 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 9JT (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | C [auth A], S [auth B] | N-phenyl-2-selanylbenzamide C13 H11 N O Se PVPUYGNPKBMXGO-UHFFFAOYSA-N |  | ||
| IOD Download:Ideal Coordinates CCD File | AA [auth B] G [auth A] H [auth A] I [auth A] J [auth A] | IODIDE ION I XMBWDFGMSWQBCA-UHFFFAOYSA-M |  | ||
| GOL Download:Ideal Coordinates CCD File | F [auth A], V [auth B] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| ZN Download:Ideal Coordinates CCD File | D [auth A], T [auth B] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| FMT Download:Ideal Coordinates CCD File | BA [auth B], M [auth A], N [auth A], O [auth A] | FORMIC ACID C H2 O2 BDAGIHXWWSANSR-UHFFFAOYSA-N |  | ||
| CL Download:Ideal Coordinates CCD File | E [auth A], U [auth B] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| NA Download:Ideal Coordinates CCD File | CA [auth B], P [auth A] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 114.887 | α = 90 |
| b = 114.887 | β = 90 |
| c = 252.82 | γ = 120 |
| Software Name | Purpose |
|---|---|
| HKL-3000 | data scaling |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| HKL-3000 | data reduction |
| HKL-3000 | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID) | United States | HHSN272201200026C |
| National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID) | United States | HHSN272201700060C |