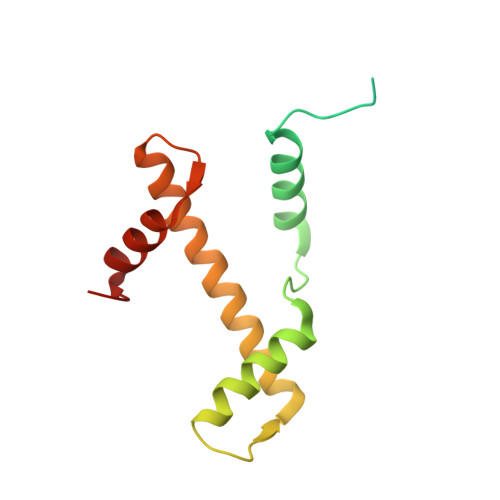
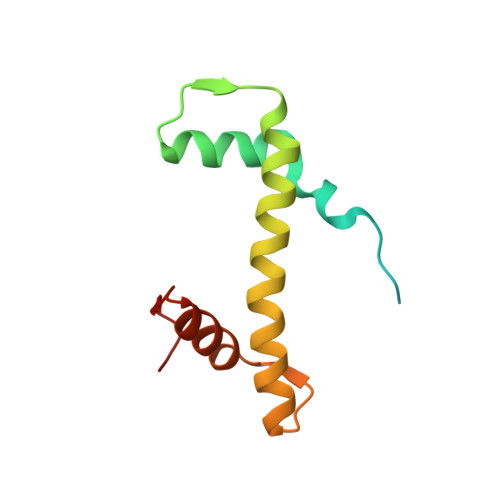
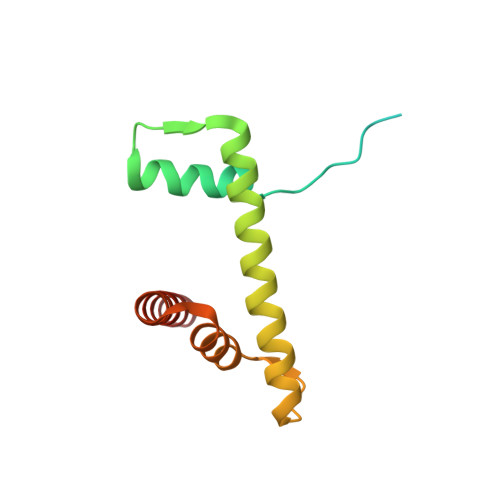
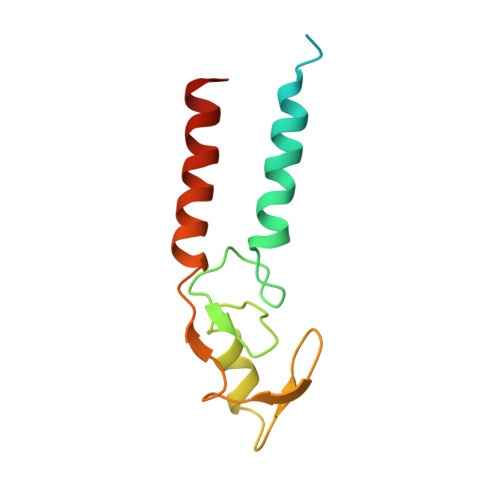
Mechanisms of BRCA1-BARD1 nucleosome recognition and ubiquitylation.
Hu, Q., Botuyan, M.V., Zhao, D., Cui, G., Mer, E., Mer, G.(2021) Nature 596: 438-443
- PubMed: 34321665 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-021-03716-8
- Primary Citation Related Structures:
7LYA, 7LYB, 7LYC - PubMed Abstract:
The BRCA1-BARD1 tumour suppressor is an E3 ubiquitin ligase necessary for the repair of DNA double-strand breaks by homologous recombination 1-10 . The BRCA1-BARD1 complex localizes to damaged chromatin after DNA replication and catalyses the ubiquitylation of histone H2A and other cellular targets 11-14 . The molecular bases for the recruitment to double-strand breaks and target recognition of BRCA1-BARD1 remain unknown. Here we use cryo-electron microscopy to show that the ankyrin repeat and tandem BRCT domains in BARD1 adopt a compact fold and bind to nucleosomal histones, DNA and monoubiquitin attached to H2A amino-terminal K13 or K15, two signals known to be specific for double-strand breaks 15,16 . We further show that RING domains 17 in BRCA1-BARD1 orient an E2 ubiquitin-conjugating enzyme atop the nucleosome in a dynamic conformation, primed for ubiquitin transfer to the flexible carboxy-terminal tails of H2A and variant H2AX. Our work reveals a regulatory crosstalk in which recognition of monoubiquitin by BRCA1-BARD1 at the N terminus of H2A blocks the formation of polyubiquitin chains and cooperatively promotes ubiquitylation at the C terminus of H2A. These findings elucidate the mechanisms of BRCA1-BARD1 chromatin recruitment and ubiquitylation specificity, highlight key functions of BARD1 in both processes and explain how BRCA1-BARD1 promotes homologous recombination by opposing the DNA repair protein 53BP1 in post-replicative chromatin 18-22 . These data provide a structural framework to evaluate BARD1 variants and help to identify mutations that drive the development of cancer.
- Department of Biochemistry and Molecular Biology, Mayo Clinic, Rochester, MN, USA.
Organizational Affiliation: