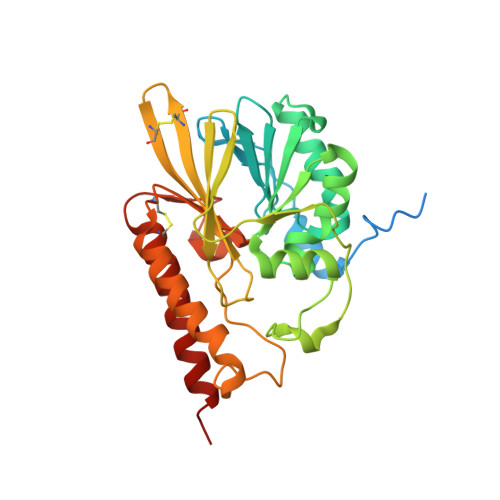
Kinetic and Structural Characterization of the First B3 Metallo-beta-Lactamase with an Active-Site Glutamic Acid.
Wilson, L.A., Knaven, E.G., Morris, M.T., Monteiro Pedroso, M., Schofield, C.J., Bruck, T.B., Boden, M., Waite, D.W., Hugenholtz, P., Guddat, L., Schenk, G.(2021) Antimicrob Agents Chemother 65: e0093621-e0093621
- PubMed: 34310207 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/AAC.00936-21
- Primary Citation Related Structures:
7LUU - PubMed Abstract:
The structural diversity in metallo-β-lactamases (MBLs), especially in the vicinity of the active site, has been a major hurdle in the development of clinically effective inhibitors. Representatives from three variants of the B3 MBL subclass, containing either the canonical HHH/DHH active-site motif (present in the majority of MBLs in this subclass) or the QHH/DHH (B3-Q) or HRH/DQK (B3-RQK) variations, were reported previously. Here, we describe the structure and kinetic properties of the first example (SIE-1) of a fourth variant containing the EHH/DHH active-site motif (B3-E). SIE-1 was identified in the hexachlorocyclohexane-degrading bacterium Sphingobium indicum, and kinetic analyses demonstrate that although it is active against a wide range of antibiotics, its efficiency is lower than that of other B3 MBLs but has increased efficiency toward cephalosporins relative to other β-lactam substrates. The overall fold of SIE-1 is characteristic of the MBLs; the notable variation is observed in the Zn1 site due to the replacement of the canonical His116 by a glutamate. The unusual preference of SIE-1 for cephalosporins and its occurrence in a widespread environmental organism suggest the scope for increased MBL-mediated β-lactam resistance. Thus, it is relevant to include SIE-1 in MBL inhibitor design studies to widen the therapeutic scope of much needed antiresistance drugs.
- School of Chemistry and Molecular Biosciences, The University of Queenslandgrid.1003.2, St. Lucia, QLD, Brisbane, Australia.
Organizational Affiliation: