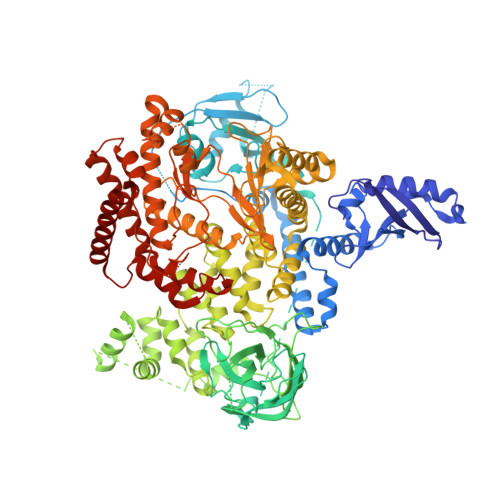
Projected Dose Optimization of Amino- and Hydroxypyrrolidine Purine PI3K delta Immunomodulators.
Methot, J.L., Zhou, H., McGowan, M.A., Anthony, N.J., Christopher, M., Garcia, Y., Achab, A., Lipford, K., Trotter, B.W., Altman, M.D., Fradera, X., Lesburg, C.A., Li, C., Alves, S., Chappell, C.P., Jain, R., Mangado, R., Pinheiro, E., Williams, S.M.G., Goldenblatt, P., Hill, A., Shaffer, L., Chen, D., Tong, V., McLeod, R.L., Lee, H.H., Yu, H., Shah, S., Katz, J.D.(2021) J Med Chem 64: 5137-5156
- PubMed: 33797901 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.1c00237
- Primary Citation Related Structures:
7LM2 - PubMed Abstract:
The approvals of idelalisib and duvelisib have validated PI3Kδ inhibitors for the treatment for hematological malignancies driven by the PI3K/AKT pathway. Our program led to the identification of structurally distinct heterocycloalkyl purine inhibitors with excellent isoform and kinome selectivity; however, they had high projected human doses. Improved ligand contacts gave potency enhancements, while replacement of metabolic liabilities led to extended half-lives in preclinical species, affording PI3Kδ inhibitors with low once-daily predicted human doses. Treatment of C57BL/6-Foxp3-GDL reporter mice with 30 and 100 mg/kg/day of 3c (MSD-496486311) led to a 70% reduction in Foxp3-expressing regulatory T cells as observed through bioluminescence imaging with luciferin, consistent with the role of PI3K/AKT signaling in Treg cell proliferation. As a model for allergic rhinitis and asthma, treatment of ovalbumin-challenged Brown Norway rats with 0.3 to 30 mg/kg/day of 3c gave a dose-dependent reduction in pulmonary bronchoalveolar lavage inflammation eosinophil cell count.
- Discovery Chemistry, Merck & Co., Inc., 33 Avenue Louis Pasteur, Boston, Massachusetts 02115 United States.
Organizational Affiliation: