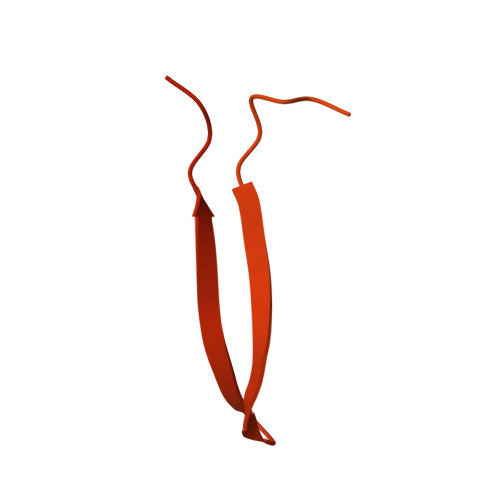
Scaffolding proteins guide the evolution of algal light harvesting antennas.
Rathbone, H.W., Michie, K.A., Landsberg, M.J., Green, B.R., Curmi, P.M.G.(2021) Nat Commun 12: 1890-1890
- PubMed: 33767155
- DOI: https://doi.org/10.1038/s41467-021-22128-w
- Primary Citation Related Structures:
7LIX, 7LIY, 7LIZ, 7LJ0 - PubMed Abstract:
Photosynthetic organisms have developed diverse antennas composed of chromophorylated proteins to increase photon capture. Cryptophyte algae acquired their photosynthetic organelles (plastids) from a red alga by secondary endosymbiosis. Cryptophytes lost the primary red algal antenna, the red algal phycobilisome, replacing it with a unique antenna composed of αβ protomers, where the β subunit originates from the red algal phycobilisome. The origin of the cryptophyte antenna, particularly the unique α subunit, is unknown. Here we show that the cryptophyte antenna evolved from a complex between a red algal scaffolding protein and phycoerythrin β. Published cryo-EM maps for two red algal phycobilisomes contain clusters of unmodelled density homologous to the cryptophyte-αβ protomer. We modelled these densities, identifying a new family of scaffolding proteins related to red algal phycobilisome linker proteins that possess multiple copies of a cryptophyte-α-like domain. These domains bind to, and stabilise, a conserved hydrophobic surface on phycoerythrin β, which is the same binding site for its primary partner in the red algal phycobilisome, phycoerythrin α. We propose that after endosymbiosis these scaffolding proteins outcompeted the primary binding partner of phycoerythrin β, resulting in the demise of the red algal phycobilisome and emergence of the cryptophyte antenna.
- School of Physics, University of New South Wales, Sydney, NSW, 2052, Australia.
Organizational Affiliation: