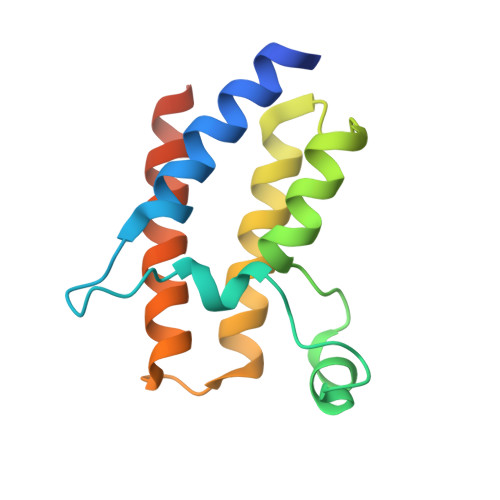
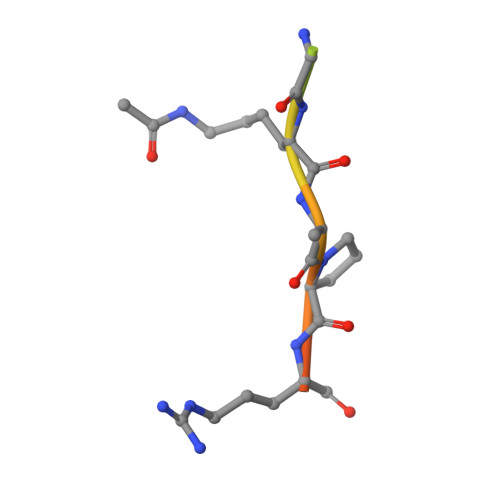
Binding specificity and function of the SWI/SNF subunit SMARCA4 bromodomain interaction with acetylated histone H3K14.
Enriquez, P., Krajewski, K., Strahl, B.D., Rothbart, S.B., Dowen, R.H., Rose, R.B.(2021) J Biological Chem 297: 101145-101145
- PubMed: 34473995 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2021.101145
- Primary Citation Related Structures:
7LHY - PubMed Abstract:
Bromodomains (BD) are conserved reader modules that bind acetylated lysine residues on histones. Although much has been learned regarding the in vitro properties of these domains, less is known about their function within chromatin complexes. SWI/SNF chromatin-remodeling complexes modulate transcription and contribute to DNA damage repair. Mutations in SWI/SNF subunits have been implicated in many cancers. Here we demonstrate that the BD of Caenorhabditis elegans SMARCA4/BRG1, a core SWI/SNF subunit, recognizes acetylated lysine 14 of histone H3 (H3K14ac), similar to its Homo sapiens ortholog. We identify the interactions of SMARCA4 with the acetylated histone peptide from a 1.29 Å-resolution crystal structure of the CeSMARCA4 BD-H3K14ac complex. Significantly, most of the SMARCA4 BD residues in contact with the histone peptide are conserved with other proteins containing family VIII bromodomains. Based on the premise that binding specificity is conserved among bromodomain orthologs, we propose that loop residues outside of the binding pocket position contact residues to recognize the H3K14ac sequence. CRISPR-Cas9-mediated mutations in the SMARCA4 BD that abolish H3K14ac binding in vitro had little or no effect on C. elegans viability or physiological function in vivo. However, combining SMARCA4 BD mutations with knockdown of the SWI/SNF accessory subunit PBRM-1 resulted in severe developmental defects in animals. In conclusion, we demonstrated an essential function for the SWI/SNF bromodomain in vivo and detected potential redundancy in epigenetic readers in regulating chromatin remodeling. These findings have implications for the development of small-molecule BD inhibitors to treat cancers and other diseases.
- Department of Molecular and Structural Biochemistry, North Carolina State University, Raleigh, North Carolina, USA.
Organizational Affiliation: