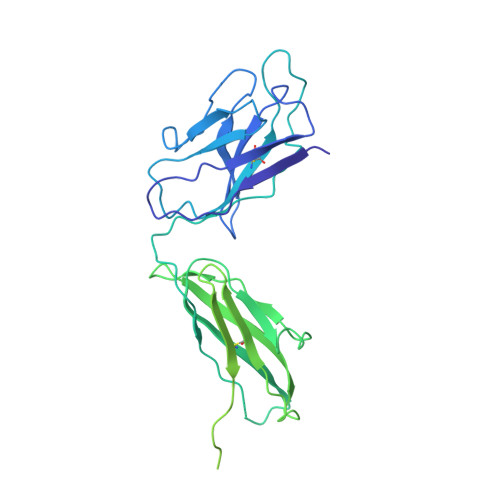
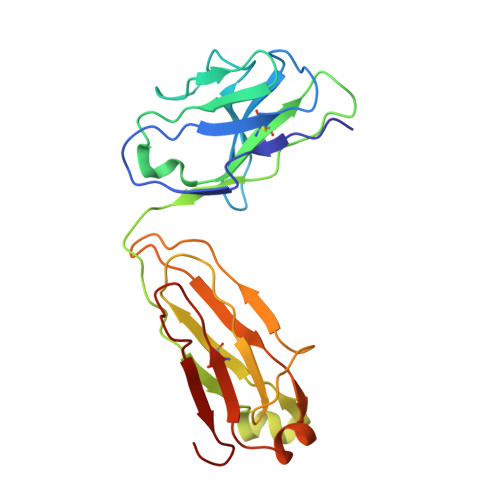
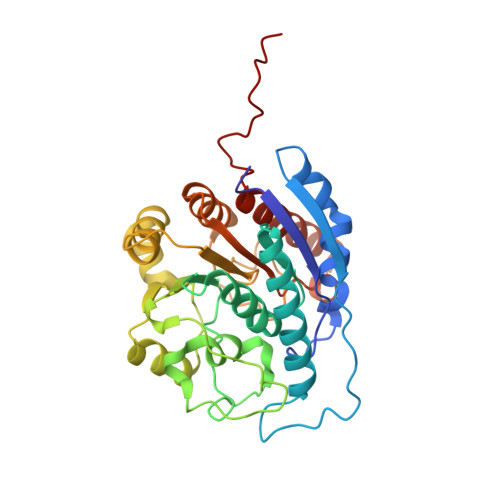
Cryo-EM structures of inhibitory antibodies complexed with arginase 1 provide insight into mechanism of action.
Palte, R.L., Juan, V., Gomez-Llorente, Y., Bailly, M.A., Chakravarthy, K., Chen, X., Cipriano, D., Fayadat-Dilman, L., Gathiaka, S., Greb, H., Hall, B., Handa, M., Hsieh, M., Kofman, E., Lin, H., Miller, J.R., Nguyen, N., O'Neil, J., Shaheen, H., Sterner, E., Strickland, C., Sun, A., Taremi, S., Scapin, G.(2021) Commun Biol 4: 927-927
- PubMed: 34326456 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-021-02444-z
- Primary Citation Related Structures:
7LEX, 7LEY, 7LEZ, 7LF0, 7LF1, 7LF2 - PubMed Abstract:
Human Arginase 1 (hArg1) is a metalloenzyme that catalyzes the hydrolysis of L-arginine to L-ornithine and urea, and modulates T-cell-mediated immune response. Arginase-targeted therapies have been pursued across several disease areas including immunology, oncology, nervous system dysfunction, and cardiovascular dysfunction and diseases. Currently, all published hArg1 inhibitors are small molecules usually less than 350 Da in size. Here we report the cryo-electron microscopy structures of potent and inhibitory anti-hArg antibodies bound to hArg1 which form distinct macromolecular complexes that are greater than 650 kDa. With local resolutions of 3.5 Å or better we unambiguously mapped epitopes and paratopes for all five antibodies and determined that the antibodies act through orthosteric and allosteric mechanisms. These hArg1:antibody complexes present an alternative mechanism to inhibit hArg1 activity and highlight the ability to utilize antibodies as probes in the discovery and development of peptide and small molecule inhibitors for enzymes in general.
- Department of Discovery Chemistry, Merck & Co., Inc., Boston, MA, USA. rachel.kubiak@merck.com.
Organizational Affiliation: