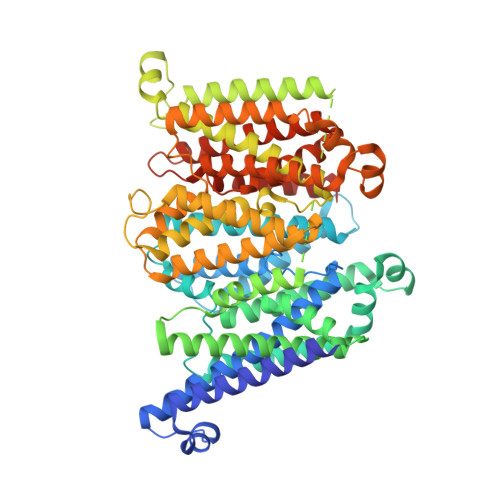
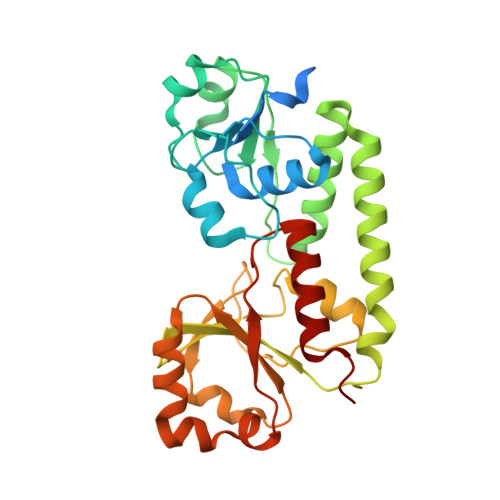
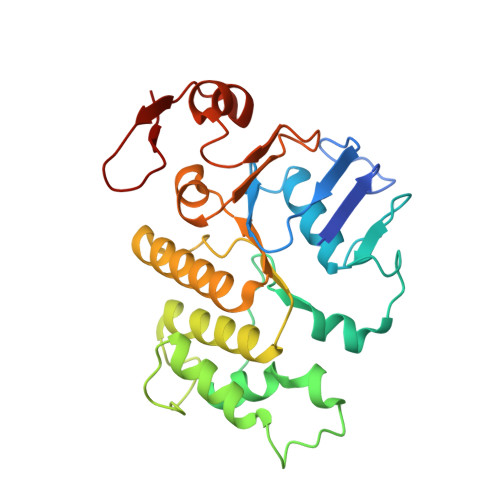
Cryo-EM reveals unique structural features of the FhuCDB Escherichia coli ferrichrome importer.
Hu, W., Zheng, H.(2021) Commun Biol 4: 1383-1383
- PubMed: 34887516
- DOI: https://doi.org/10.1038/s42003-021-02916-2
- Primary Citation Related Structures:
7LB8 - PubMed Abstract:
As one of the most elegant biological processes developed in bacteria, the siderophore-mediated iron uptake demands the action of specific ATP-binding cassette (ABC) importers. Although extensive studies have been done on various ABC importers, the molecular basis of these iron-chelated-siderophore importers are still not fully understood. Here, we report the structure of a ferrichrome importer FhuCDB from Escherichia coli at 3.4 Å resolution determined by cryo electron microscopy. The structure revealed a monomeric membrane subunit of FhuB with a substrate translocation pathway in the middle. In the pathway, there were unique arrangements of residues, especially layers of methionines. Important residues found in the structure were interrogated by mutagenesis and functional studies. Surprisingly, the importer's ATPase activity was decreased upon FhuD binding, which deviated from the current understanding about bacterial ABC importers. In summary, to the best of our knowledge, these studies not only reveal a new structural twist in the type II ABC importer subfamily, but also provide biological insights in the transport of iron-chelated siderophores.
- Department of Biochemistry and Molecular Genetics, University of Colorado Anschutz Medical Campus, School of Medicine, Aurora, USA.
Organizational Affiliation: