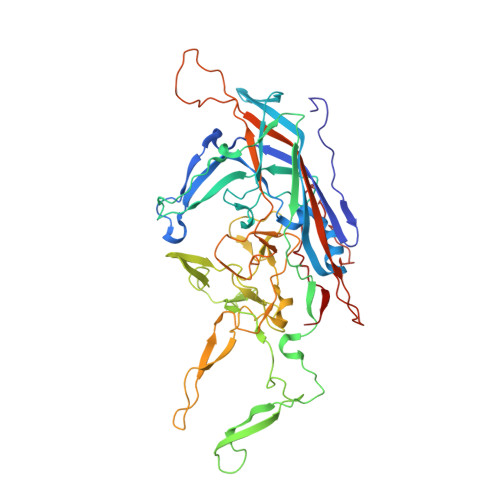
Completion of the AAV Structural Atlas: Serotype Capsid Structures Reveals Clade-Specific Features.
Mietzsch, M., Jose, A., Chipman, P., Bhattacharya, N., Daneshparvar, N., McKenna, R., Agbandje-McKenna, M.(2021) Viruses 13
- PubMed: 33450892 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/v13010101
- Primary Citation Related Structures:
7L5Q, 7L5U, 7L6A, 7L6B, 7L6E, 7L6F, 7L6H, 7L6I - PubMed Abstract:
The capsid structures of most Adeno-associated virus (AAV) serotypes, already assigned to an antigenic clade, have been previously determined. This study reports the remaining capsid structures of AAV7, AAV11, AAV12, and AAV13 determined by cryo-electron microscopy and three-dimensional image reconstruction to 2.96, 2.86, 2.54, and 2.76 Å resolution, respectively. These structures complete the structural atlas of the AAV serotype capsids. AAV7 represents the first clade D capsid structure; AAV11 and AAV12 are of a currently unassigned clade that would include AAV4; and AAV13 represents the first AAV2-AAV3 hybrid clade C capsid structure. These newly determined capsid structures all exhibit the AAV capsid features including 5-fold channels, 3-fold protrusions, 2-fold depressions, and a nucleotide binding pocket with an ordered nucleotide in genome-containing capsids. However, these structures have viral proteins that display clade-specific loop conformations. This structural characterization completes our three-dimensional library of the current AAV serotypes to provide an atlas of surface loop configurations compatible with capsid assembly and amenable for future vector engineering efforts. Derived vectors could improve gene delivery success with respect to specific tissue targeting, transduction efficiency, antigenicity or receptor retargeting.
- Department of Biochemistry and Molecular Biology, Center for Structural Biology, McKnight Brain Institute, College of Medicine, University of Florida, Gainesville, FL 32610, USA.
Organizational Affiliation: