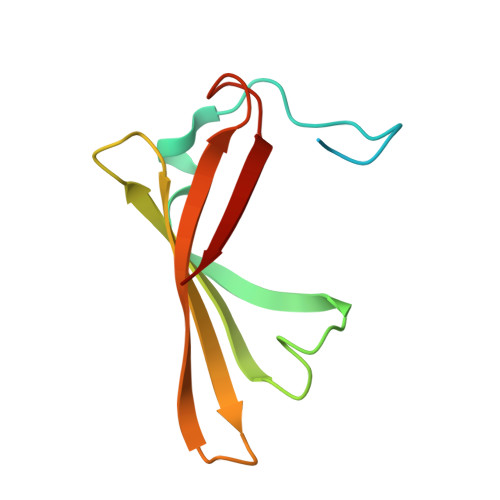
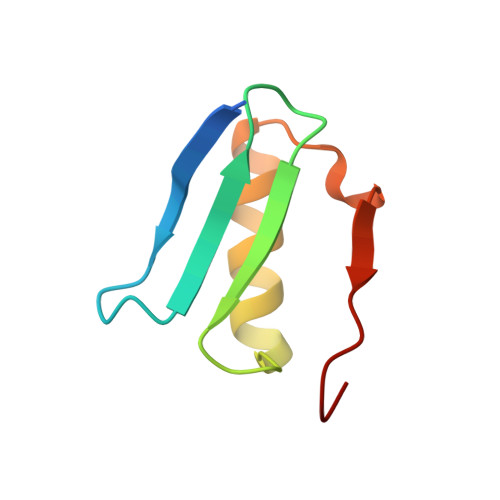
Computational redesign of a fluorogen activating protein with Rosetta.
Bozhanova, N.G., Harp, J.M., Bender, B.J., Gavrikov, A.S., Gorbachev, D.A., Baranov, M.S., Mercado, C.B., Zhang, X., Lukyanov, K.A., Mishin, A.S., Meiler, J.(2021) PLoS Comput Biol 17: e1009555-e1009555
- PubMed: 34748541 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pcbi.1009555
- Primary Citation Related Structures:
7L5K, 7L5L, 7L5M - PubMed Abstract:
The use of unnatural fluorogenic molecules widely expands the pallet of available genetically encoded fluorescent imaging tools through the design of fluorogen activating proteins (FAPs). While there is already a handful of such probes available, each of them went through laborious cycles of in vitro screening and selection. Computational modeling approaches are evolving incredibly fast right now and are demonstrating great results in many applications, including de novo protein design. It suggests that the easier task of fine-tuning the fluorogen-binding properties of an already functional protein in silico should be readily achievable. To test this hypothesis, we used Rosetta for computational ligand docking followed by protein binding pocket redesign to further improve the previously described FAP DiB1 that is capable of binding to a BODIPY-like dye M739. Despite an inaccurate initial docking of the chromophore, the incorporated mutations nevertheless improved multiple photophysical parameters as well as the overall performance of the tag. The designed protein, DiB-RM, shows higher brightness, localization precision, and apparent photostability in protein-PAINT super-resolution imaging compared to its parental variant DiB1. Moreover, DiB-RM can be cleaved to obtain an efficient split system with enhanced performance compared to a parental DiB-split system. The possible reasons for the inaccurate ligand binding pose prediction and its consequence on the outcome of the design experiment are further discussed.
- Department of Chemistry and Center for Structural Biology, Vanderbilt University, Nashville, Tennessee, United States of America.
Organizational Affiliation: