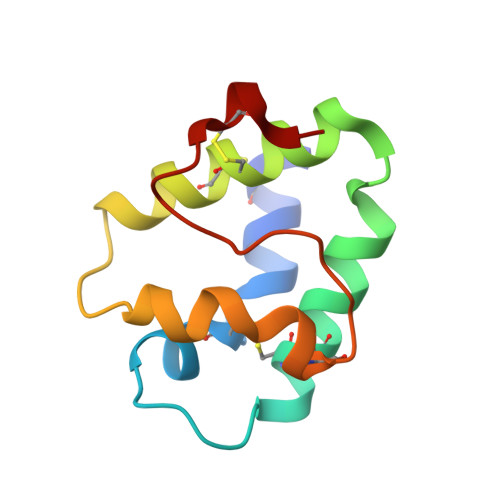
Structural Characterization of Act c 10.0101 and Pun g 1.0101-Allergens from the Non-Specific Lipid Transfer Protein Family.
O'Malley, A., Pote, S., Giangrieco, I., Tuppo, L., Gawlicka-Chruszcz, A., Kowal, K., Ciardiello, M.A., Chruszcz, M.(2021) Molecules 26
- PubMed: 33419110
- DOI: https://doi.org/10.3390/molecules26020256
- Primary Citation Related Structures:
7KSB, 7KSC - PubMed Abstract:
(1) Background: Non-specific lipid transfer proteins (nsLTPs), which belong to the prolamin superfamily, are potent allergens. While the biological role of LTPs is still not well understood, it is known that these proteins bind lipids. Allergen nsLTPs are characterized by significant stability and resistance to digestion. (2) Methods: nsLTPs from gold kiwifruit (Act c 10.0101) and pomegranate (Pun g 1.0101) were isolated from their natural sources and structurally characterized using X-ray crystallography (3) Results: Both proteins crystallized and their crystal structures were determined. The proteins have a very similar overall fold with characteristic compact, mainly α-helical structures. The C-terminal sequence of Act c 10.0101 was updated based on our structural and mass spectrometry analysis. Information on proteins' sequences and structures was used to estimate the risk of cross-reactive reactions between Act c 10.0101 or Pun g 1.0101 and other allergens from this family of proteins. (4) Conclusions: Structural studies indicate a conformational flexibility of allergens from the nsLTP family and suggest that immunoglobulin E binding to some surface regions of these allergens may depend on ligand binding. Both Act c 10.0101 and Pun g 1.0101 are likely to be involved in cross-reactive reactions involving other proteins from the nsLTP family.
- Department of Chemistry and Biochemistry, University of South Carolina, Columbia, SC 29208, USA.
Organizational Affiliation: