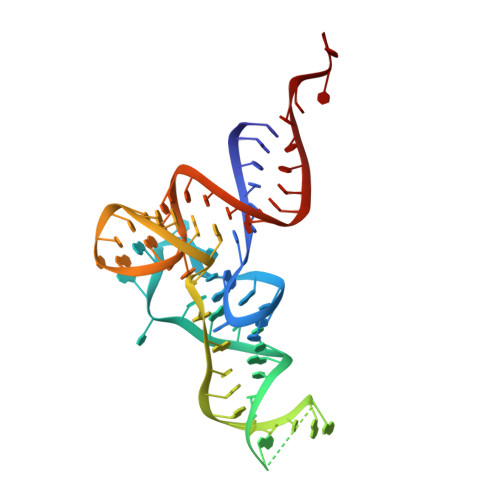
A substrate binding model for the KEOPS tRNA modifying complex.
Beenstock, J., Ona, S.M., Porat, J., Orlicky, S., Wan, L.C.K., Ceccarelli, D.F., Maisonneuve, P., Szilard, R.K., Yin, Z., Setiaputra, D., Mao, D.Y.L., Khan, M., Raval, S., Schriemer, D.C., Bayfield, M.A., Durocher, D., Sicheri, F.(2020) Nat Commun 11: 6233-6233
- PubMed: 33277478 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-19990-5
- Primary Citation Related Structures:
7KJT, 7KJU - PubMed Abstract:
The KEOPS complex, which is conserved across archaea and eukaryotes, is composed of four core subunits; Pcc1, Kae1, Bud32 and Cgi121. KEOPS is crucial for the fitness of all organisms examined. In humans, pathogenic mutations in KEOPS genes lead to Galloway-Mowat syndrome, an autosomal-recessive disease causing childhood lethality. Kae1 catalyzes the universal and essential tRNA modification N 6 -threonylcarbamoyl adenosine, but the precise roles of all other KEOPS subunits remain an enigma. Here we show using structure-guided studies that Cgi121 recruits tRNA to KEOPS by binding to its 3' CCA tail. A composite model of KEOPS bound to tRNA reveals that all KEOPS subunits form an extended tRNA-binding surface that we have validated in vitro and in vivo to mediate the interaction with the tRNA substrate and its modification. These findings provide a framework for understanding the inner workings of KEOPS and delineate why all KEOPS subunits are essential.
- The Lunenfeld-Tanenbaum Research Institute, Mount Sinai Hospital, Toronto, ON, Canada.
Organizational Affiliation: