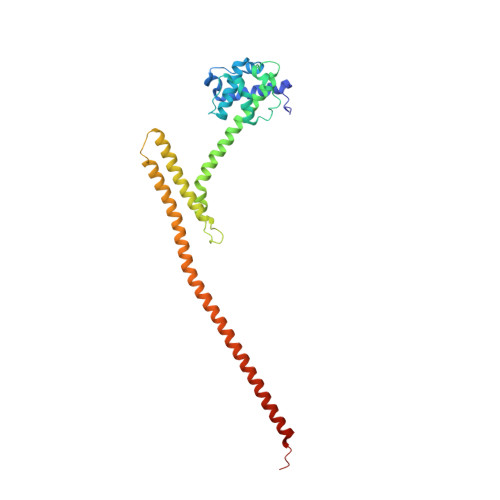
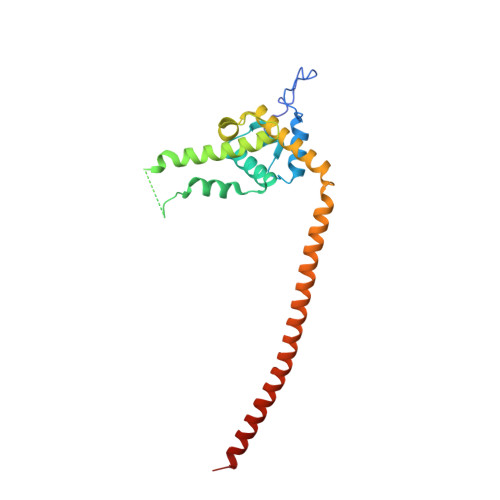
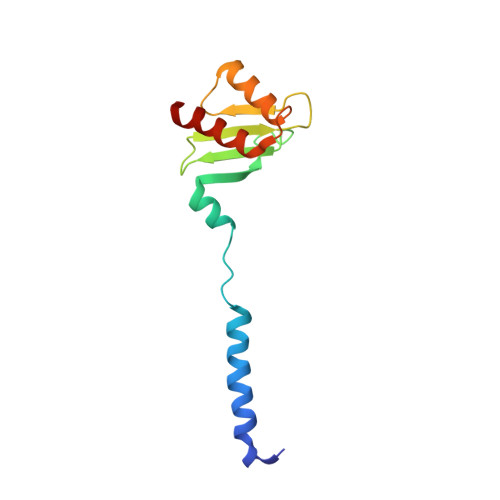
Structural basis of Stu2 recruitment to yeast kinetochores.
Zahm, J.A., Stewart, M.G., Carrier, J.S., Harrison, S.C., Miller, M.P.(2021) Elife 10
- PubMed: 33591274 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.65389
- Primary Citation Related Structures:
7KDF - PubMed Abstract:
Chromosome segregation during cell division requires engagement of kinetochores of sister chromatids with microtubules emanating from opposite poles. As the corresponding microtubules shorten, these 'bioriented' sister kinetochores experience tension-dependent stabilization of microtubule attachments. The yeast XMAP215 family member and microtubule polymerase, Stu2, associates with kinetochores and contributes to tension-dependent stabilization in vitro. We show here that a C-terminal segment of Stu2 binds the four-way junction of the Ndc80 complex (Ndc80c) and that residues conserved both in yeast Stu2 orthologs and in their metazoan counterparts make specific contacts with Ndc80 and Spc24. Mutations that perturb this interaction prevent association of Stu2 with kinetochores, impair cell viability, produce biorientation defects, and delay cell cycle progression. Ectopic tethering of the mutant Stu2 species to the Ndc80c junction restores wild-type function in vivo. These findings show that the role of Stu2 in tension-sensing depends on its association with kinetochores by binding with Ndc80c.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, and Howard Hughes Medical Institute, Boston, United States.
Organizational Affiliation: