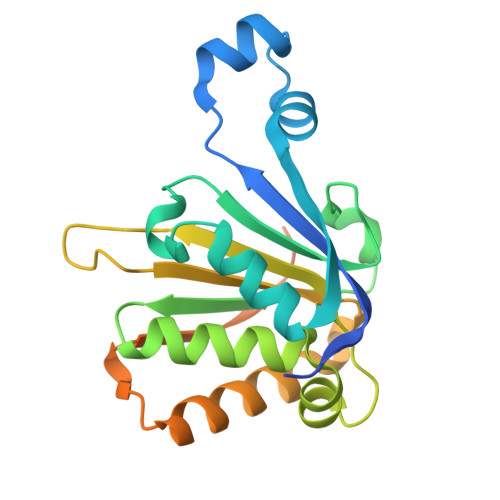
The translation initiation factor EIF4E5 from Leishmania: crystal structure and interacting partners.
de Lima, G.B., de Lima Cavalcanti, T.Y.V., de Brito, A.N.A.L.M., de Assis, L.A., Andrade-Vieira, R.P., Freire, E.R., da Silva Assuncao, T.R., de Souza Reis, C.R., Zanchin, N.I.T., Guimaraes, B.G., de-Melo-Neto, O.P.(2021) RNA Biol 18: 2433-2449
- PubMed: 33945405 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1080/15476286.2021.1918919
- Primary Citation Related Structures:
7KCJ - PubMed Abstract:
The mRNA cap-binding protein, eIF4E, mediates the recognition of the mRNA 5' end and, as part of the heterotrimeric eIF4F complex, facilitates the recruitment of the ribosomal subunits to initiate eukaryotic translation. Various regulatory events involving eIF4E and a second eIF4F subunit, eIF4G, are required for proper control of translation initiation. In pathogenic trypanosomatids, six eIF4Es and five eIF4Gs have been described, several forming different eIF4F-like complexes with yet unresolved roles. EIF4E5 is one of the least known of the trypanosomatid eIF4Es and has not been characterized in Leishmania species. Here, we used immunoprecipitation assays, combined with mass-spectrometry, to identify major EIF4E5 interacting proteins in L. infantum . A constitutively expressed, HA-tagged, EIF4E5 co-precipitated mainly with EIF4G1 and binding partners previously described in Trypanosoma brucei , EIF4G1-IP, RBP43 and the 14-3-3 proteins. In contrast, no clear co-precipitation with EIF4G2, also previously reported, was observed. EIF4E5 also co-precipitated with protein kinases, possibly associated with cell-cycle regulation, selected RNA binding proteins and histones. Phosphorylated residues were identified and mapped to the Leishmania -specific C-terminal end. Mutagenesis of the tryptophan residue (W53) postulated to mediate interactions with protein partners or of a neighbouring tryptophan conserved in Leishmania (W45) did not substantially impair the identified interactions. Finally, the crystal structure of Leishmania EIF4E5 evidences remarkable differences in the eIF4G interfacing region, when compared with human eIF4E-1 and with its Trypanosoma orthologue. Mapping of its C-terminal end near the cap-binding site also imply relevant differences in cap-binding function and/or regulation.
- Departamento de Microbiologia, Instituto Aggeu Magalhães, FIOCRUZ-PE, Av. Moraes Rego s/n, Recife-PE, Brazil.
Organizational Affiliation: