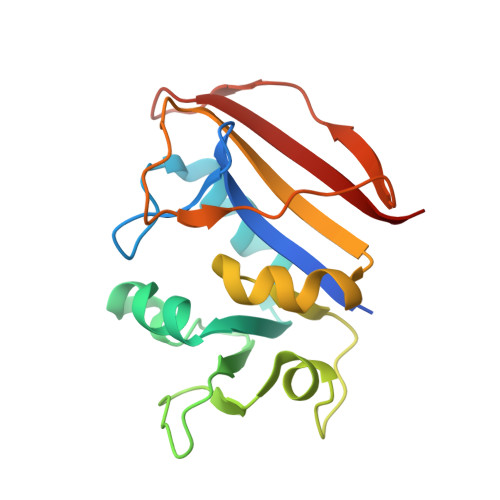
Crystal Structure of Dihydrofolate reductase (DHFR) from Mycobacterium ulcerans Agy99 in complex with NADP and inhibitor SDDC-0001575
Abendroth, J., Dranow, D.M., Santhakumar, V., Walpole, C., Lorimer, D.D., Horanyi, P.S., Edwards, T.E.To be published.