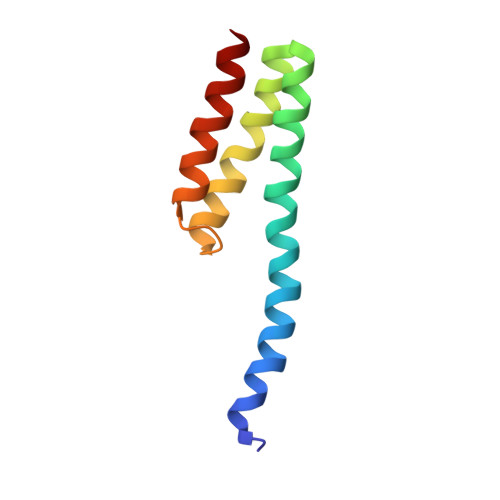
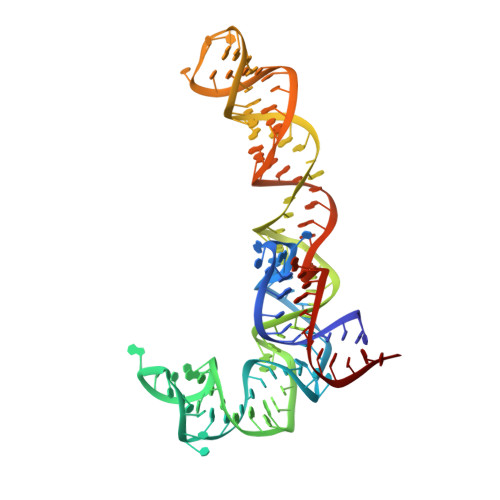
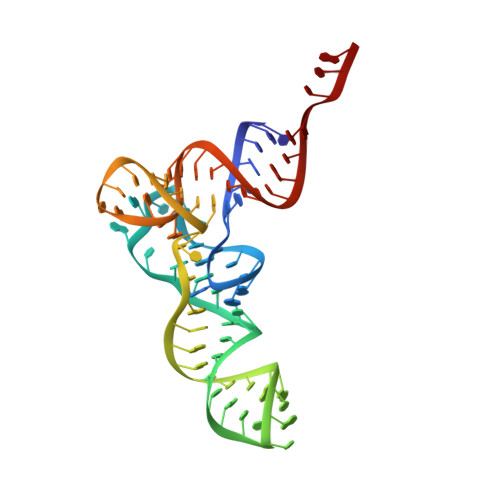
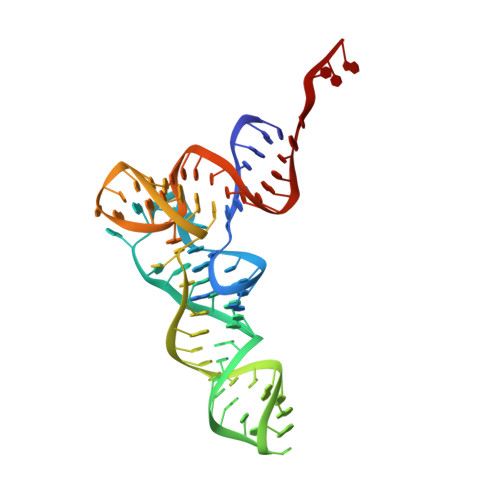
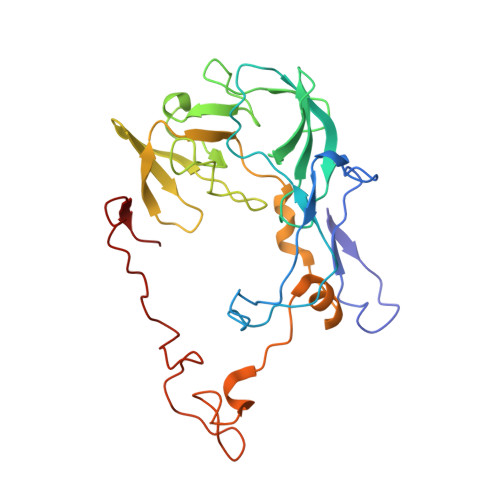
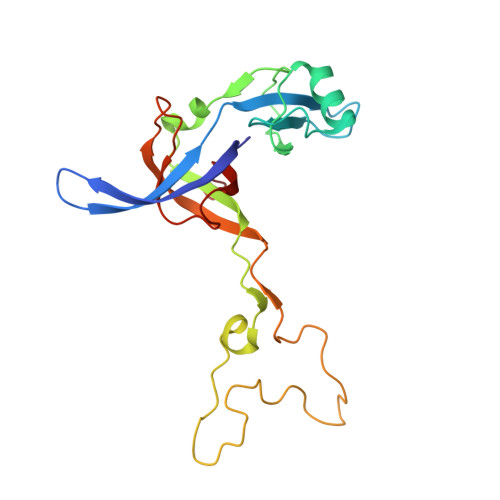
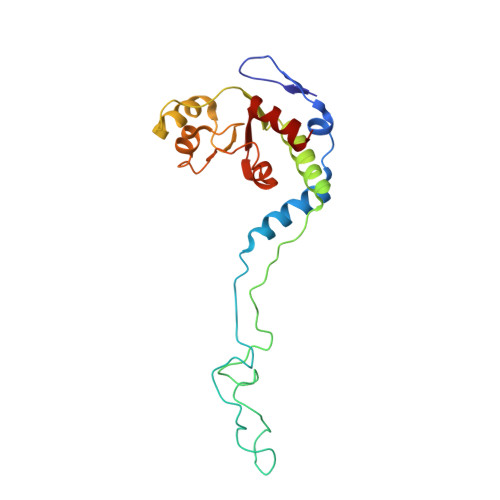
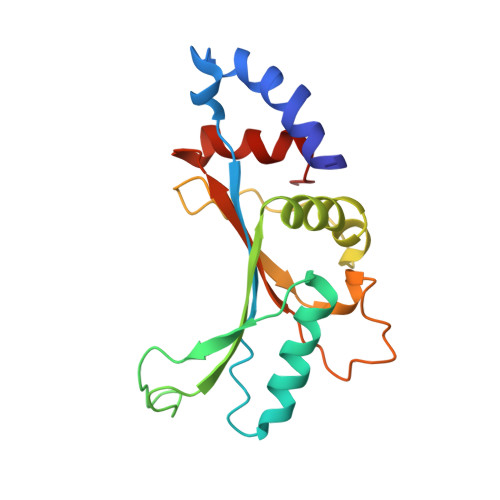
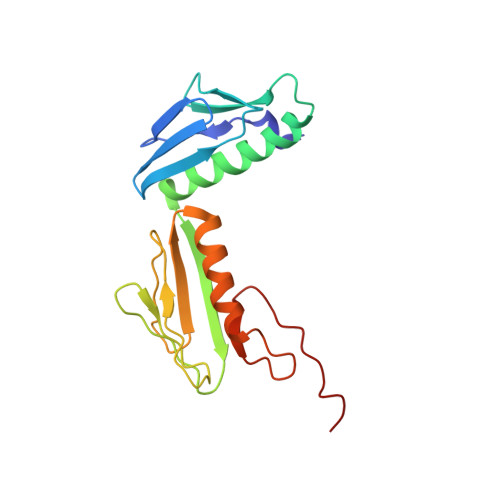
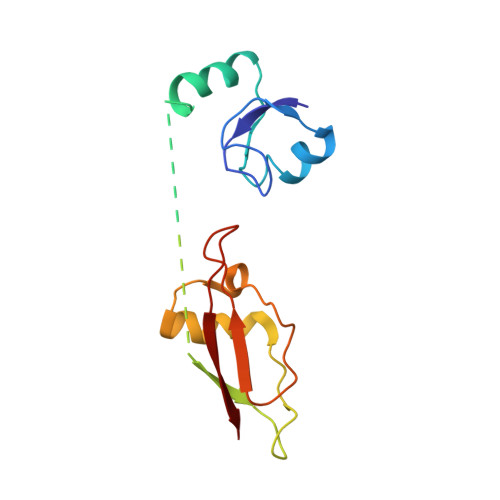
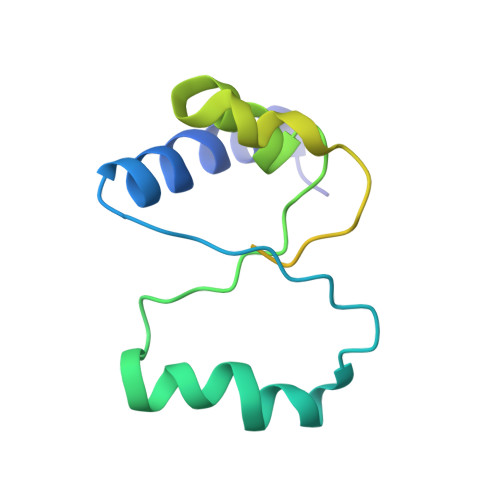
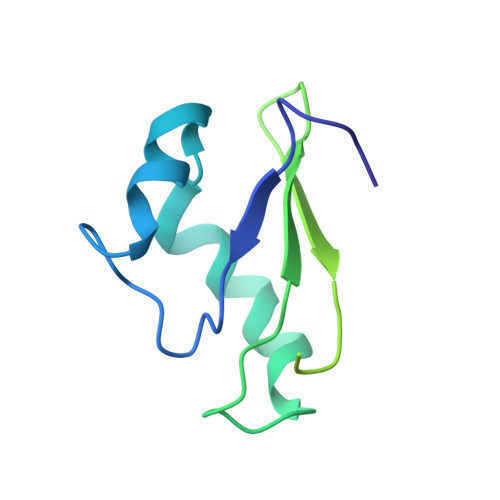
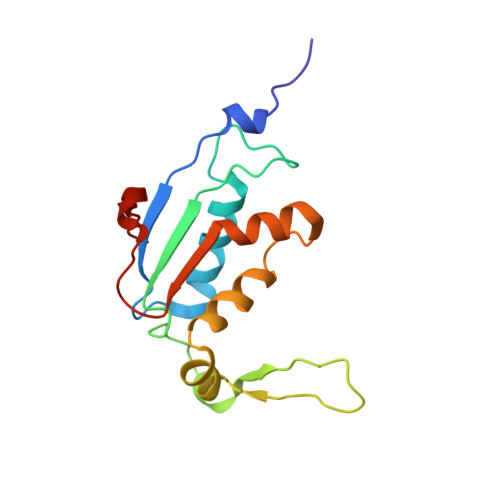
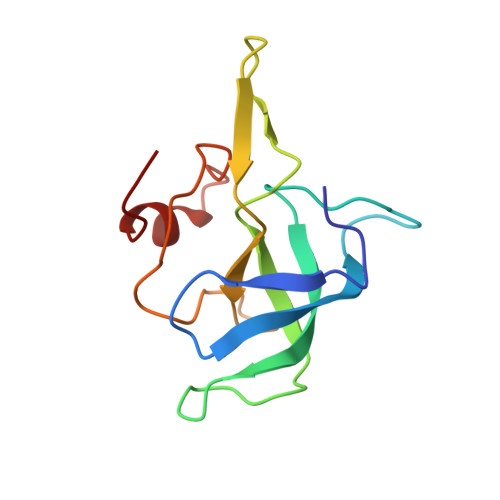
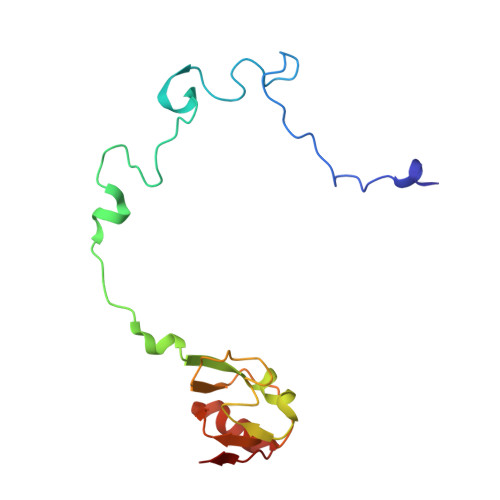
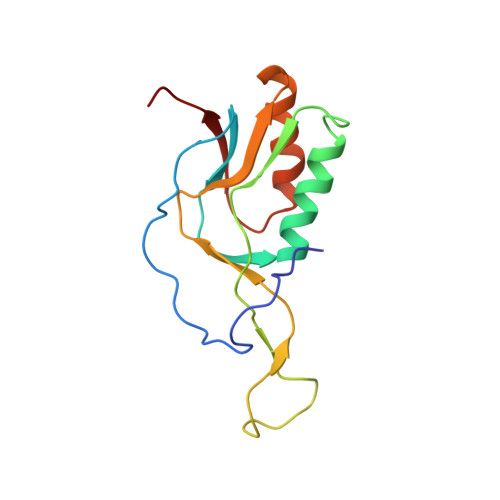
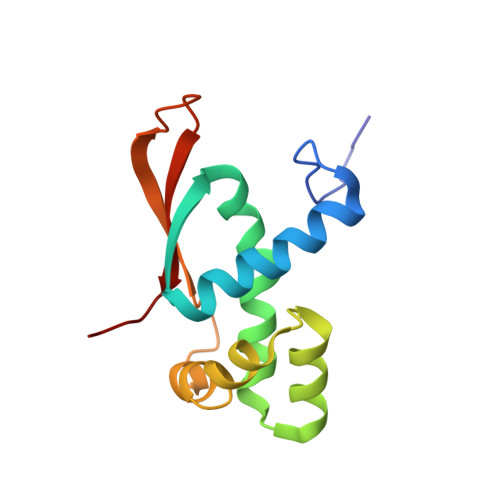
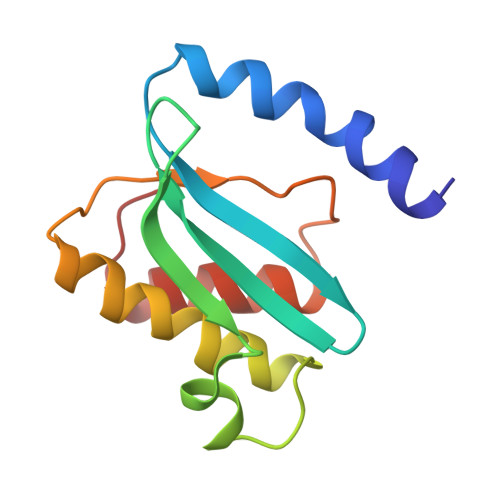
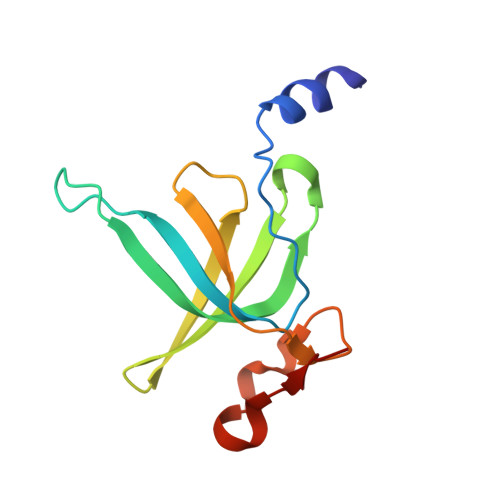
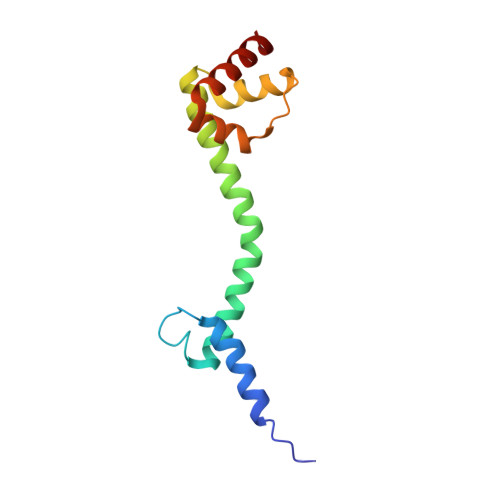
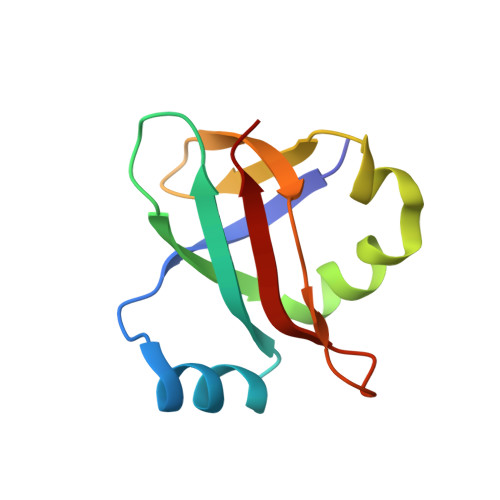
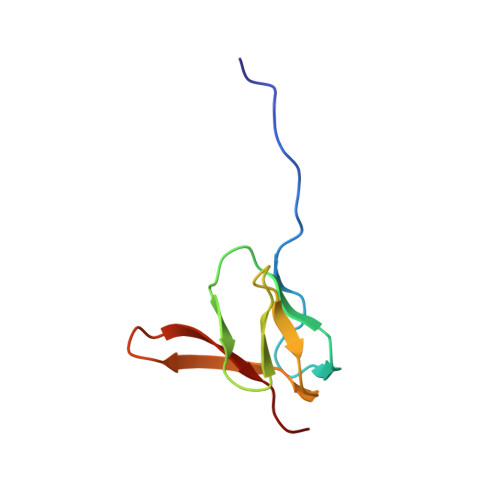
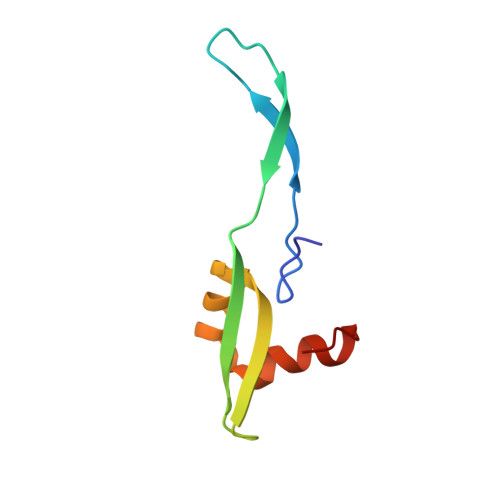
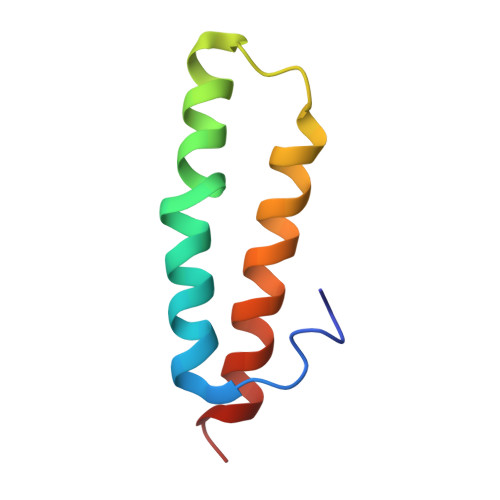
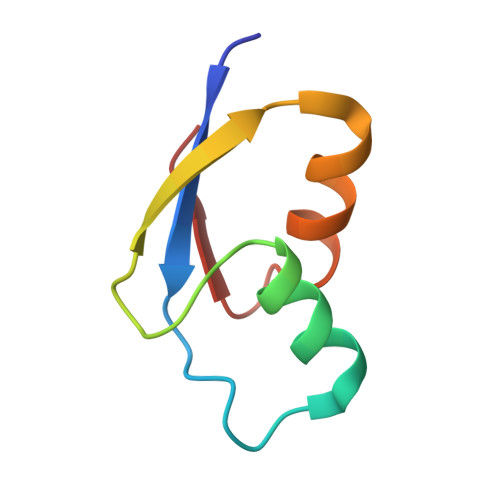
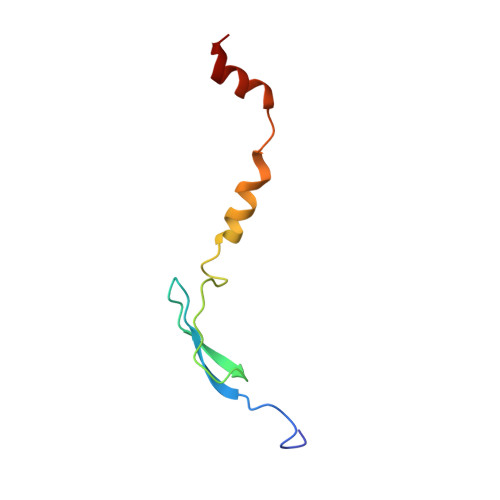
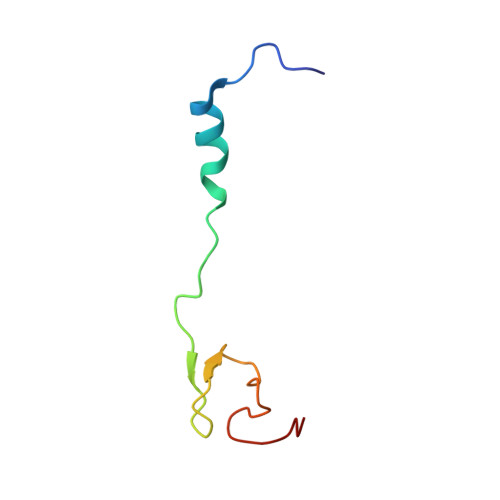
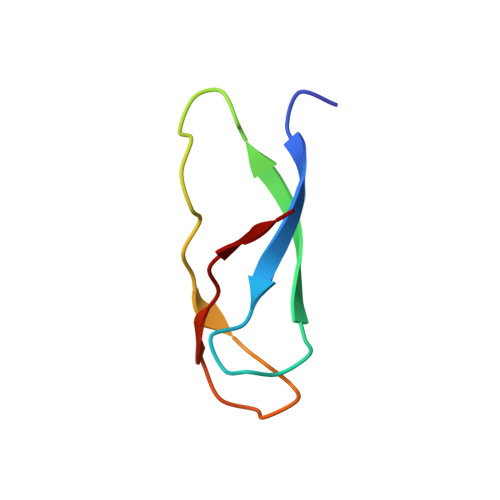
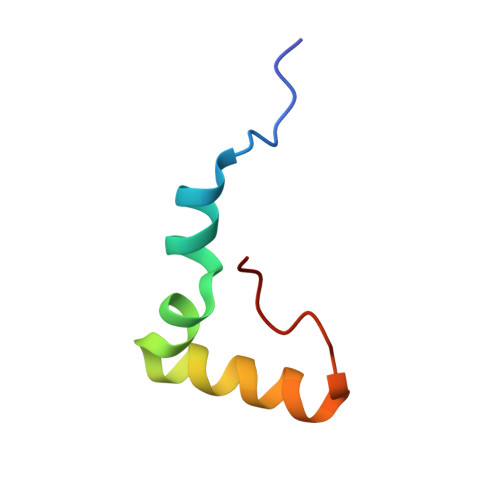
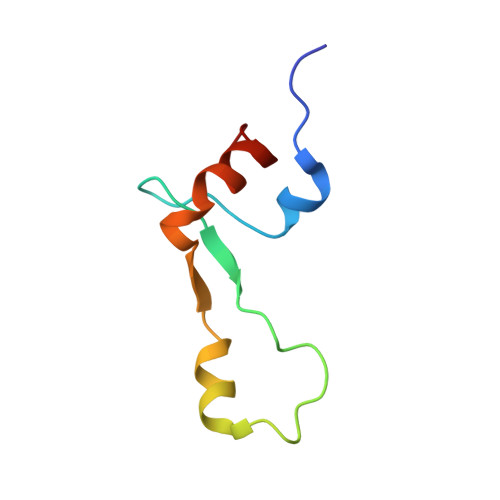
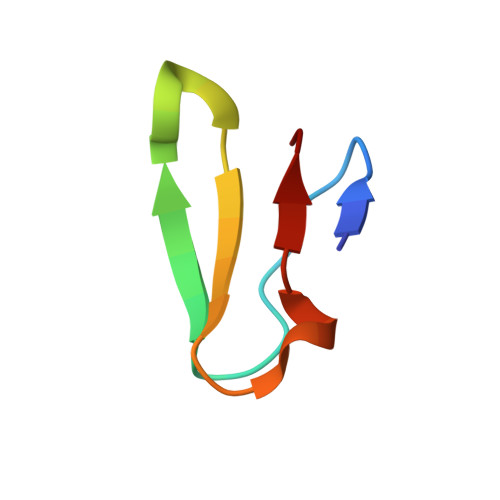
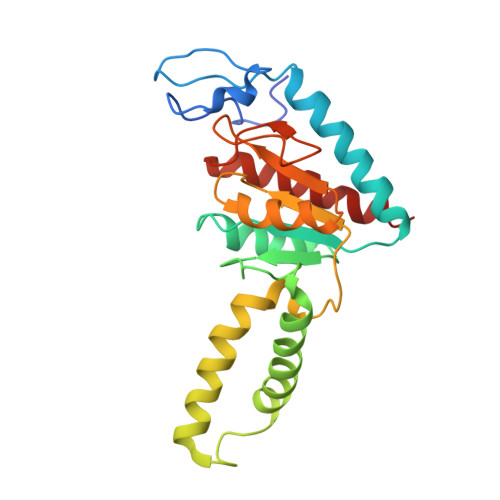
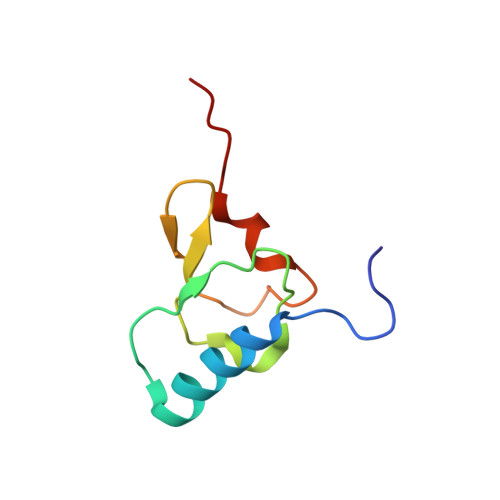
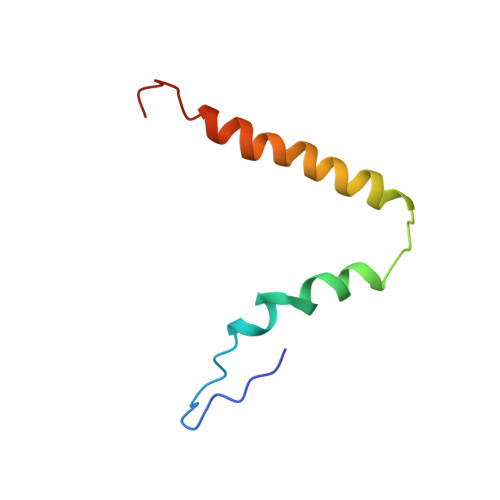
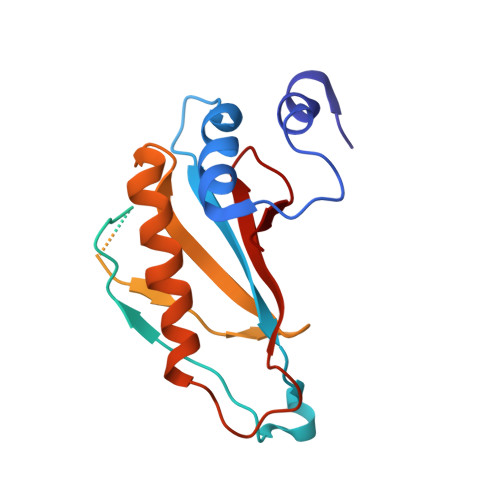
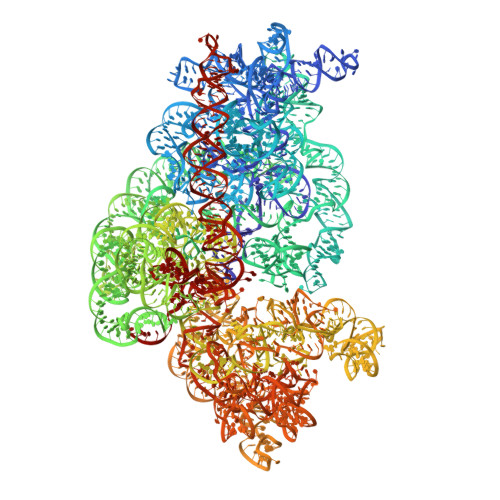
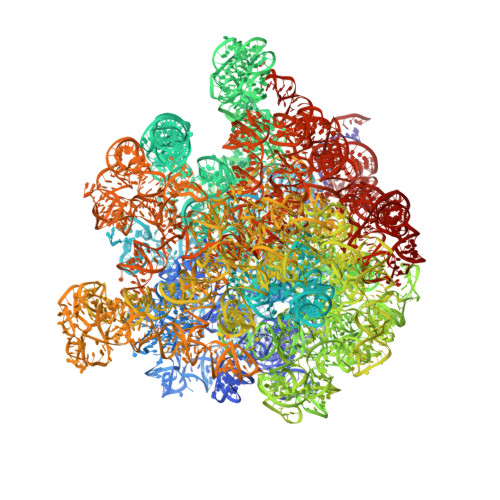
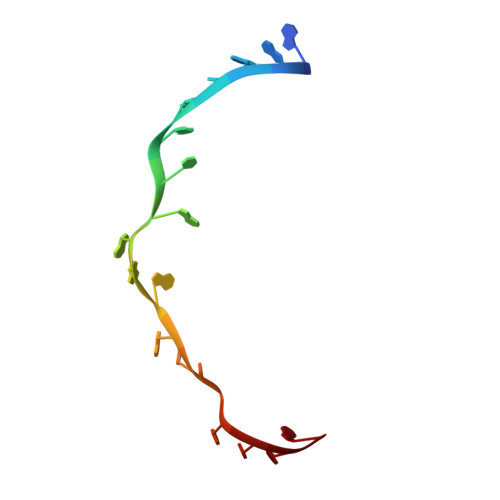
Structural basis for +1 ribosomal frameshifting during EF-G-catalyzed translocation.
Demo, G., Gamper, H.B., Loveland, A.B., Masuda, I., Carbone, C.E., Svidritskiy, E., Hou, Y.M., Korostelev, A.A.(2021) Nat Commun 12: 4644-4644
- PubMed: 34330903 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-24911-1
- Primary Citation Related Structures:
7K50, 7K51, 7K52, 7K53, 7K54, 7K55, 7LV0 - PubMed Abstract:
Frameshifting of mRNA during translation provides a strategy to expand the coding repertoire of cells and viruses. How and where in the elongation cycle +1-frameshifting occurs remains poorly understood. We describe seven ~3.5-Å-resolution cryo-EM structures of 70S ribosome complexes, allowing visualization of elongation and translocation by the GTPase elongation factor G (EF-G). Four structures with a + 1-frameshifting-prone mRNA reveal that frameshifting takes place during translocation of tRNA and mRNA. Prior to EF-G binding, the pre-translocation complex features an in-frame tRNA-mRNA pairing in the A site. In the partially translocated structure with EF-G•GDPCP, the tRNA shifts to the +1-frame near the P site, rendering the freed mRNA base to bulge between the P and E sites and to stack on the 16S rRNA nucleotide G926. The ribosome remains frameshifted in the nearly post-translocation state. Our findings demonstrate that the ribosome and EF-G cooperate to induce +1 frameshifting during tRNA-mRNA translocation.
- RNA Therapeutics Institute, Department of Biochemistry and Molecular Pharmacology, UMass Medical School, Worcester, MA, USA.
Organizational Affiliation: