Structural basis for voltage-sensor trapping of the cardiac sodium channel by a deathstalker scorpion toxin.
Jiang, D., Tonggu, L., Gamal El-Din, T.M., Banh, R., Pomes, R., Zheng, N., Catterall, W.A.(2021) Nat Commun 12: 128-128
- PubMed: 33397917 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-20078-3
- Primary Citation Related Structures:
7K18 - PubMed Abstract:
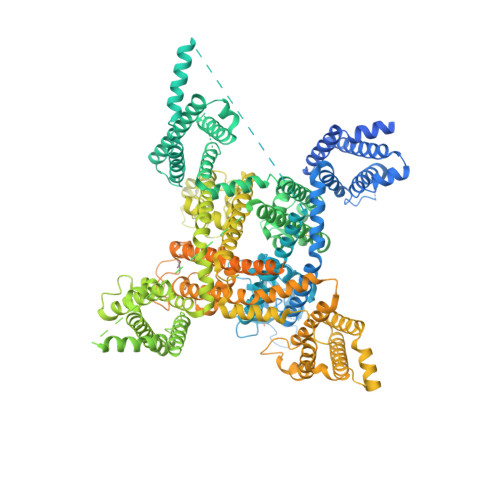
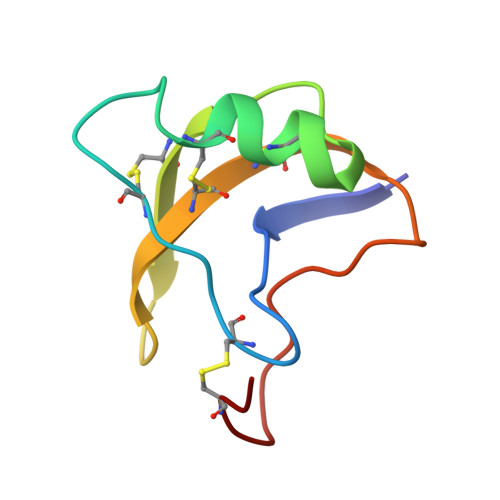
Voltage-gated sodium (Na V ) channels initiate action potentials in excitable cells, and their function is altered by potent gating-modifier toxins. The α-toxin LqhIII from the deathstalker scorpion inhibits fast inactivation of cardiac Na V 1.5 channels with IC 50 = 11.4 nM. Here we reveal the structure of LqhIII bound to Na V 1.5 at 3.3 Å resolution by cryo-EM. LqhIII anchors on top of voltage-sensing domain IV, wedged between the S1-S2 and S3-S4 linkers, which traps the gating charges of the S4 segment in a unique intermediate-activated state stabilized by four ion-pairs. This conformational change is propagated inward to weaken binding of the fast inactivation gate and favor opening the activation gate. However, these changes do not permit Na + permeation, revealing why LqhIII slows inactivation of Na V channels but does not open them. Our results provide important insights into the structural basis for gating-modifier toxin binding, voltage-sensor trapping, and fast inactivation of Na V channels.
- Department of Pharmacology, University of Washington, Seattle, WA, 98195, USA.
Organizational Affiliation: