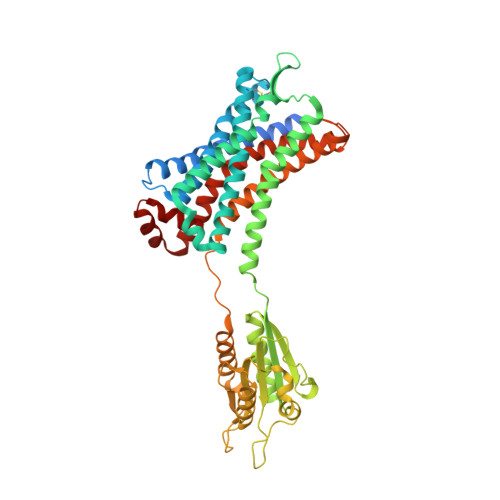
Structural insights on ligand recognition at the human leukotriene B4 receptor 1.
Michaelian, N., Sadybekov, A., Besserer-Offroy, E., Han, G.W., Krishnamurthy, H., Zamlynny, B.A., Fradera, X., Siliphaivanh, P., Presland, J., Spencer, K.B., Soisson, S.M., Popov, P., Sarret, P., Katritch, V., Cherezov, V.(2021) Nat Commun 12: 2971-2971
- PubMed: 34016973 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-23149-1
- Primary Citation Related Structures:
7K15 - PubMed Abstract:
The leukotriene B4 receptor 1 (BLT1) regulates the recruitment and chemotaxis of different cell types and plays a role in the pathophysiology of infectious, allergic, metabolic, and tumorigenic human diseases. Here we present a crystal structure of human BLT1 (hBLT1) in complex with a selective antagonist MK-D-046, developed for the treatment of type 2 diabetes and other inflammatory conditions. Comprehensive analysis of the structure and structure-activity relationship data, reinforced by site-directed mutagenesis and docking studies, reveals molecular determinants of ligand binding and selectivity toward different BLT receptor subtypes and across species. The structure helps to identify a putative membrane-buried ligand access channel as well as potential receptor binding modes of endogenous agonists. These structural insights of hBLT1 enrich our understanding of its ligand recognition and open up future avenues in structure-based drug design.
- Bridge Institute, USC Michelson Center for Convergent Bioscience, University of Southern California, Los Angeles, CA, USA.
Organizational Affiliation: