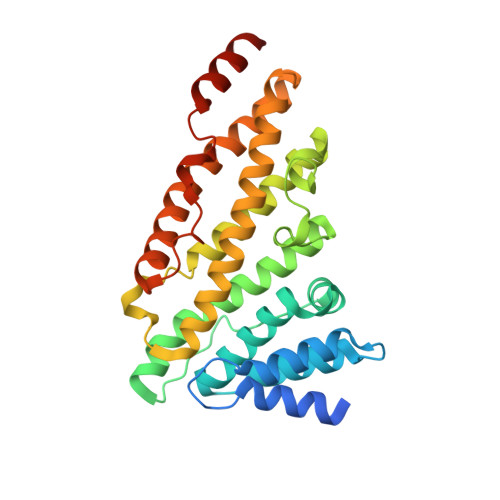
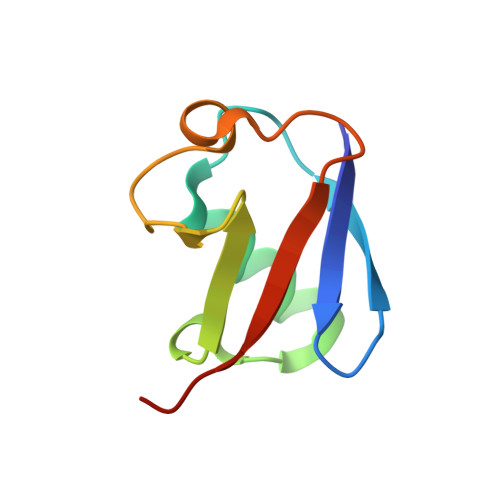
ANTH domains within CALM, HIP1R, and Sla2 recognize ubiquitin internalization signals.
Pashkova, N., Gakhar, L., Yu, L., Schnicker, N.J., Minard, A.Y., Winistorfer, S., Johnson, I.E., Piper, R.C.(2021) Elife 10
- PubMed: 34821552 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.72583
- Primary Citation Related Structures:
7JXV - PubMed Abstract:
Attachment of ubiquitin (Ub) to cell surface proteins serves as a signal for internalization via clathrin-mediated endocytosis (CME). How ubiquitinated membrane proteins engage the internalization apparatus remains unclear. The internalization apparatus contains proteins such as Epsin and Eps15, which bind Ub, potentially acting as adaptors for Ub-based internalization signals. Here, we show that additional components of the endocytic machinery including CALM, HIP1R, and Sla2 bind Ub via their N-terminal ANTH domain, a domain belonging to the superfamily of ENTH and VHS domains. Structural studies revealed that Ub binds with µM affinity to a unique C-terminal region within the ANTH domain not found in ENTH domains. Functional studies showed that combined loss of Ub-binding by ANTH-domain proteins and other Ub-binding domains within the yeast internalization apparatus caused defects in the Ub-dependent internalization of the GPCR Ste2 that was engineered to rely exclusively on Ub as an internalization signal. In contrast, these mutations had no effect on the internalization of Ste2 engineered to use an alternate Ub-independent internalization signal. These studies define new components of the internalization machinery that work collectively with Epsin and Eps15 to specify recognition of Ub as an internalization signal.
- Department of Molecular Physiology and Biophysics, University of Iowa, Iowa City, United States.
Organizational Affiliation: