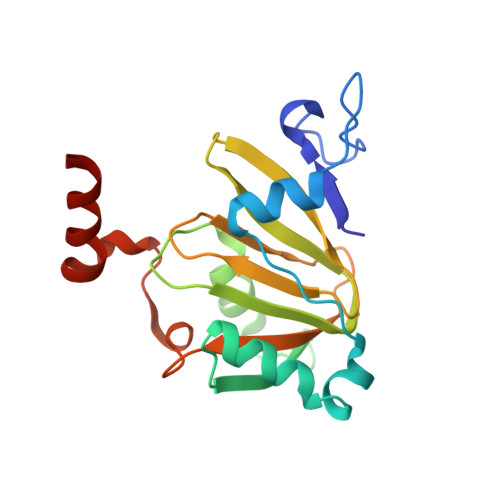
A Model for the Solution Structure of Human Fe(II)-Bound Acireductone Dioxygenase and Interactions with the Regulatory Domain of Matrix Metalloproteinase I (MMP-I).
Liu, X., Garber, A., Ryan, J., Deshpande, A., Ringe, D., Pochapsky, T.C.(2020) Biochemistry 59: 4238-4249
- PubMed: 33135413 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.0c00724
- Primary Citation Related Structures:
7JXG - PubMed Abstract:
The metalloenzyme acireductone dioxygenase (ARD) shows metal-dependent physical and enzymatic activities depending upon the metal bound in the active site. The Fe(II)-bound enzyme catalyzes the penultimate step of the methionine salvage pathway, converting 1,2-dihydroxy-5-(methylthio)pent-1-en-3-one (acireductone) into formate and the ketoacid precursor of methionine, 2-keto-4-thiomethyl-2-oxobutanoate, using O 2 as the oxidant. If Ni(II) is bound, an off-pathway shunt occurs, producing 3-methylthiopropionate, formate, and carbon monoxide from the same acireductone substrate. The solution structure of the Fe(II)-bound human enzyme, HsARD, is described and compared with the structures of Ni-bound forms of the closely related mouse enzyme, MmARD. Potential rationales for the different reactivities of the two isoforms are discussed. The human enzyme has been found to regulate the activity of matrix metalloproteinase I (MMP-I), which is involved in tumor metastasis, by binding the cytoplasmic transmembrane tail peptide of MMP-I. Nuclear magnetic resonance titration of HsARD with the MMP-I tail peptide permits identification of the peptide binding site on HsARD, a cleft anterior to the metal binding site adjacent to a dynamic proline-rich loop.
- Department of Chemistry, Brandeis University, 415 South Street, Waltham, Massachusetts 02454-9110, United States.
Organizational Affiliation: