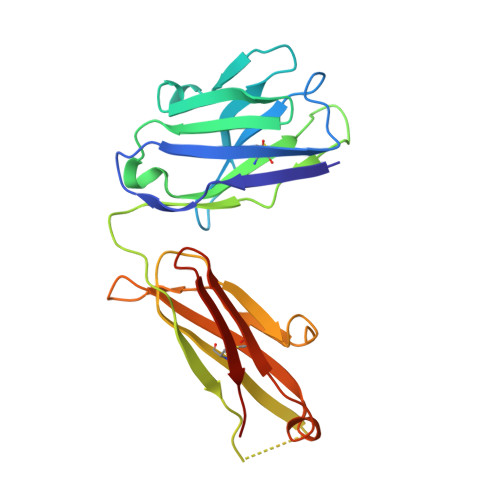
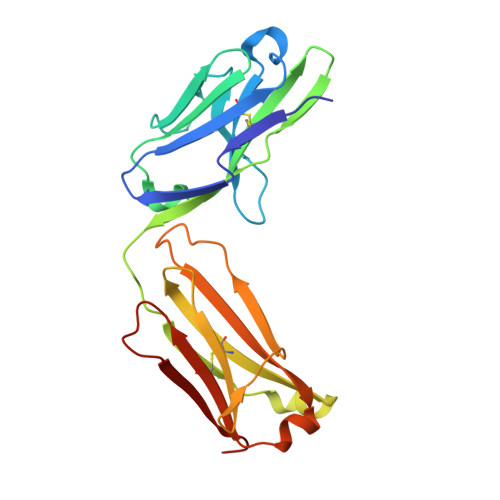
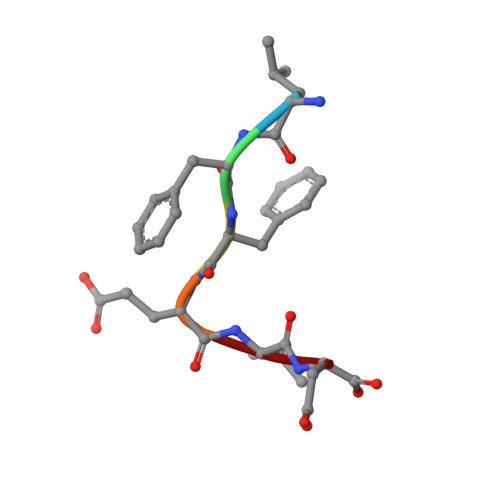
Discovery of a caspase cleavage motif antibody reveals insights into noncanonical inflammasome function.
Davies, C.W., Stowe, I., Phung, Q.T., Ho, H., Bakalarski, C.E., Gupta, A., Zhang, Y., Lill, J.R., Payandeh, J., Kayagaki, N., Koerber, J.T.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 33723046 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2018024118
- Primary Citation Related Structures:
7JWP, 7JWQ - PubMed Abstract:
Inflammasomes sense a number of pathogen and host damage signals to initiate a signaling cascade that triggers inflammatory cell death, termed pyroptosis. The inflammatory caspases (1/4/5/11) are the key effectors of this process through cleavage and activation of the pore-forming protein gasdermin D. Caspase-1 also activates proinflammatory interleukins, IL-1β and IL-18, via proteolysis. However, compared to the well-studied apoptotic caspases, the identity of substrates and therefore biological functions of the inflammatory caspases remain limited. Here, we construct, validate, and apply an antibody toolset for direct detection of neo-C termini generated by inflammatory caspase proteolysis. By combining rabbit immune phage display with a set of degenerate and defined target peptides, we discovered two monoclonal antibodies that bind peptides with a similar degenerate recognition motif as the inflammatory caspases without recognizing the canonical apoptotic caspase recognition motif. Crystal structure analyses revealed the molecular basis of this strong yet paradoxical degenerate mode of peptide recognition. One antibody selectively immunoprecipitated cleaved forms of known and unknown inflammatory caspase substrates, allowing the identification of over 300 putative substrates of the caspase-4 noncanonical inflammasome, including caspase-7. This dataset will provide a path toward developing blood-based biomarkers of inflammasome activation. Overall, our study establishes tools to discover and detect inflammatory caspase substrates and functions, provides a workflow for designing antibody reagents to study cell signaling, and extends the growing evidence of biological cross talk between the apoptotic and inflammatory caspases.
- Department of Antibody Engineering, Genentech, Inc., South San Francisco, CA 94080.
Organizational Affiliation: