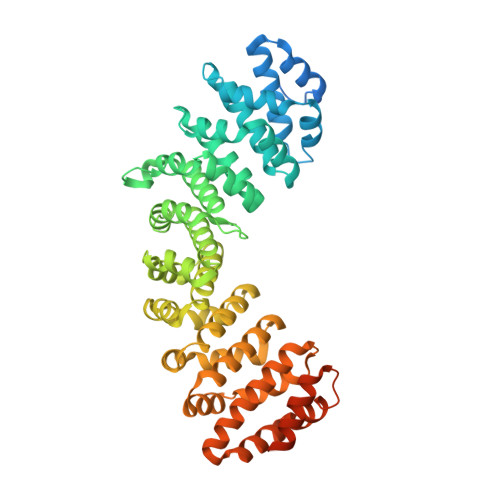
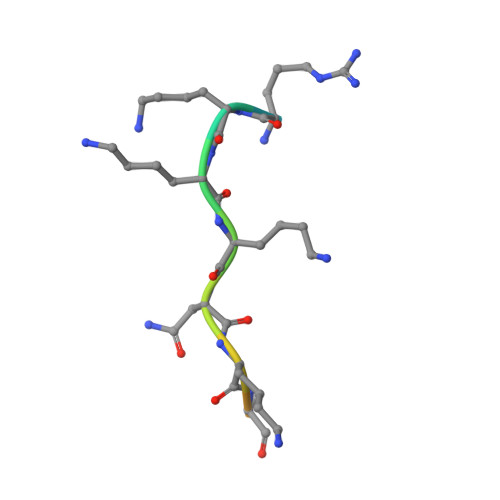
Targeting novel LSD1-dependent ACE2 demethylation domains inhibits SARS-CoV-2 replication.
Tu, W.J., McCuaig, R.D., Melino, M., Rawle, D.J., Le, T.T., Yan, K., Suhrbier, A., Johnston, R.L., Koufariotis, L.T., Waddell, N., Cross, E.M., Tsimbalyuk, S., Bain, A., Ahern, E., Collinson, N., Phipps, S., Forwood, J.K., Seddiki, N., Rao, S.(2021) Cell Discov 7: 37-37
- PubMed: 34031383 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-021-00279-w
- Primary Citation Related Structures:
7JVO - PubMed Abstract:
Treatment options for COVID-19 remain limited, especially during the early or asymptomatic phase. Here, we report a novel SARS-CoV-2 viral replication mechanism mediated by interactions between ACE2 and the epigenetic eraser enzyme LSD1, and its interplay with the nuclear shuttling importin pathway. Recent studies have shown a critical role for the importin pathway in SARS-CoV-2 infection, and many RNA viruses hijack this axis to re-direct host cell transcription. LSD1 colocalized with ACE2 at the cell surface to maintain demethylated SARS-CoV-2 spike receptor-binding domain lysine 31 to promote virus-ACE2 interactions. Two newly developed peptide inhibitors competitively inhibited virus-ACE2 interactions, and demethylase access to significantly inhibit viral replication. Similar to some other predominantly plasma membrane proteins, ACE2 had a novel nuclear function: its cytoplasmic domain harbors a nuclear shuttling domain, which when demethylated by LSD1 promoted importin-α-dependent nuclear ACE2 entry following infection to regulate active transcription. A novel, cell permeable ACE2 peptide inhibitor prevented ACE2 nuclear entry, significantly inhibiting viral replication in SARS-CoV-2-infected cell lines, outperforming other LSD1 inhibitors. These data raise the prospect of post-exposure prophylaxis for SARS-CoV-2, either through repurposed LSD1 inhibitors or new, nuclear-specific ACE2 inhibitors.
- Gene Regulation and Translational Medicine Laboratory, QIMR Berghofer Medical Research Institute, Brisbane, QLD, Australia.
Organizational Affiliation: