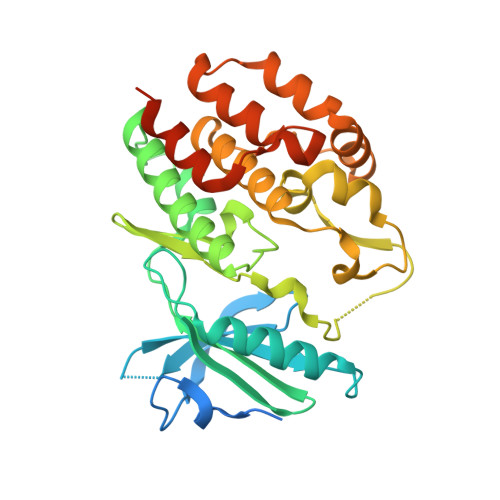
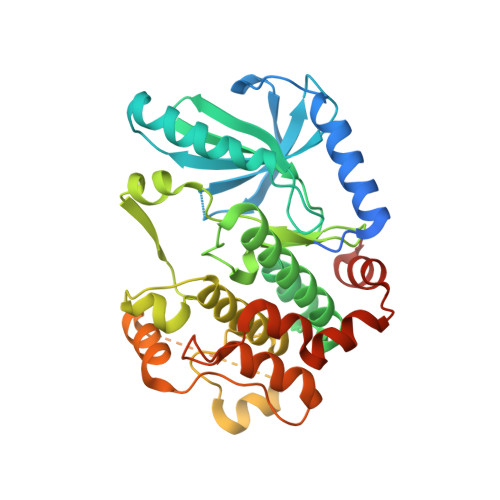
Structural basis for the action of the drug trametinib at KSR-bound MEK.
Khan, Z.M., Real, A.M., Marsiglia, W.M., Chow, A., Duffy, M.E., Yerabolu, J.R., Scopton, A.P., Dar, A.C.(2020) Nature 588: 509-514
- PubMed: 32927473 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-020-2760-4
- Primary Citation Related Structures:
7JUQ, 7JUR, 7JUS, 7JUT, 7JUU, 7JUV, 7JUW, 7JUX, 7JUY, 7JUZ, 7JV0, 7JV1 - PubMed Abstract:
The MAPK/ERK kinase MEK is a shared effector of the frequent cancer drivers KRAS and BRAF that has long been pursued as a drug target in oncology 1 , and more recently in immunotherapy 2,3 and ageing 4 . However, many MEK inhibitors are limited owing to on-target toxicities 5-7 and drug resistance 8-10 . Accordingly, a molecular understanding of the structure and function of MEK within physiological complexes could provide a template for the design of safer and more effective therapies. Here we report X-ray crystal structures of MEK bound to the scaffold KSR (kinase suppressor of RAS) with various MEK inhibitors, including the clinical drug trametinib. The structures reveal an unexpected mode of binding in which trametinib directly engages KSR at the MEK interface. In the bound complex, KSR remodels the prototypical allosteric pocket of the MEK inhibitor, thereby affecting binding and kinetics, including the drug-residence time. Moreover, trametinib binds KSR-MEK but disrupts the related RAF-MEK complex through a mechanism that exploits evolutionarily conserved interface residues that distinguish these sub-complexes. On the basis of these insights, we created trametiglue, which limits adaptive resistance to MEK inhibition by enhancing interfacial binding. Our results reveal the plasticity of an interface pocket within MEK sub-complexes and have implications for the design of next-generation drugs that target the RAS pathway.
- Department of Oncological Sciences, The Tisch Cancer Institute, Icahn School of Medicine at Mount Sinai, New York, NY, USA.
Organizational Affiliation: