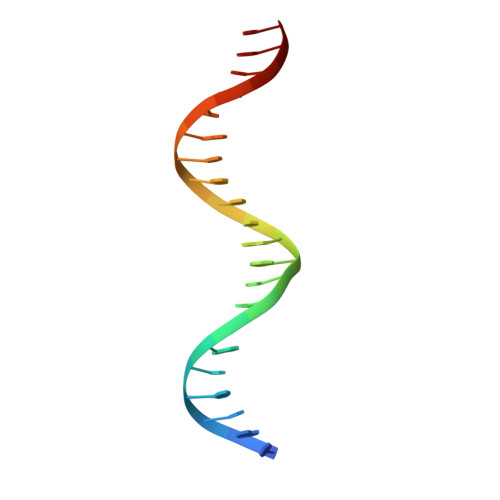
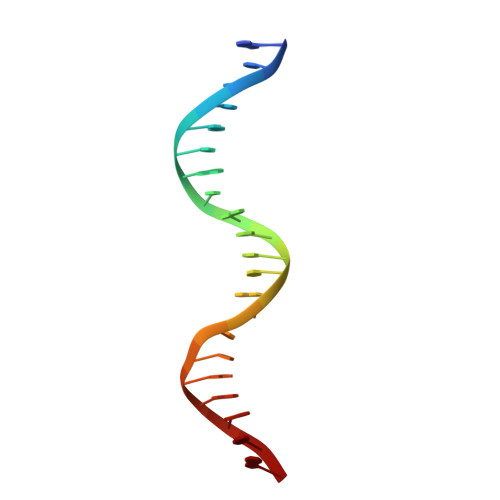
Structures of Neisseria gonorrhoeae MtrR-operator complexes reveal molecular mechanisms of DNA recognition and antibiotic resistance-conferring clinical mutations.
Beggs, G.A., Ayala, J.C., Kavanaugh, L.G., Read, T.D., Hooks, G.M., Schumacher, M.A., Shafer, W.M., Brennan, R.G.(2021) Nucleic Acids Res 49: 4155-4170
- PubMed: 33784401 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkab213
- Primary Citation Related Structures:
7JNP, 7JU3 - PubMed Abstract:
Mutations within the mtrR gene are commonly found amongst multidrug resistant clinical isolates of Neisseria gonorrhoeae, which has been labelled a superbug by the Centers for Disease Control and Prevention. These mutations appear to contribute to antibiotic resistance by interfering with the ability of MtrR to bind to and repress expression of its target genes, which include the mtrCDE multidrug efflux transporter genes and the rpoH oxidative stress response sigma factor gene. However, the DNA-recognition mechanism of MtrR and the consensus sequence within these operators to which MtrR binds has remained unknown. In this work, we report the crystal structures of MtrR bound to the mtrCDE and rpoH operators, which reveal a conserved, but degenerate, DNA consensus binding site 5'-MCRTRCRN4YGYAYGK-3'. We complement our structural data with a comprehensive mutational analysis of key MtrR-DNA contacts to reveal their importance for MtrR-DNA binding both in vitro and in vivo. Furthermore, we model and generate common clinical mutations of MtrR to provide plausible biochemical explanations for the contribution of these mutations to multidrug resistance in N. gonorrhoeae. Collectively, our findings unveil key biological mechanisms underlying the global stress responses of N. gonorrhoeae.
- Department of Biochemistry, Duke University School of Medicine, Durham, NC 27710, USA.
Organizational Affiliation: