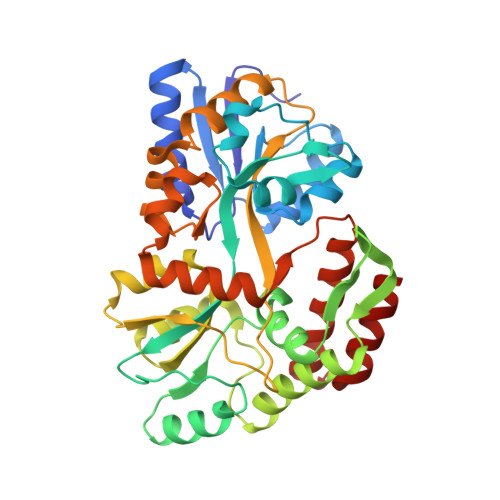
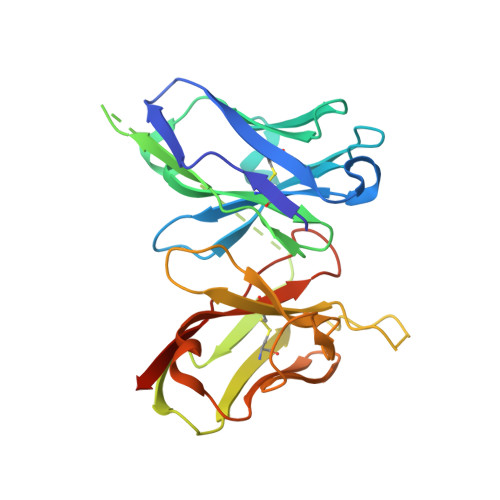
A useful epitope tag derived from maltose binding protein.
Lenon, M., Ke, N., Ren, G., Meuser, M.E., Loll, P.J., Riggs, P., Berkmen, M.(2021) Protein Sci 30: 1235-1246
- PubMed: 33896065 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.4088
- Primary Citation Related Structures:
7JTR - PubMed Abstract:
Maltose binding protein (MBP) is used in recombinant protein expression as an affinity and solubility tag. The monoclonal antibody B48 binds MBP tightly and has no cross-reactivity to other proteins in an Escherichia coli lysate. This high level of specificity suggested that MBP contains an epitope that could prove useful as a purification and visualization tag for proteins expressed in E. coli. To discover the MBP epitope, a co-crystal structure was determined for MBP bound to its antibody and four amino acids of MBP were identified as critical for the binding interaction. Fusions of various fragments of MBP to the glutathione S-transferase protein were engineered in order to identify the smallest fragment still recognized by the α-MBP antibody. Stabilization of the epitope via mutational engineering resulted in a minimized 14 amino-acid tag.
- Protein Expression and Modification, New England Biolabs, Ipswich, USA.
Organizational Affiliation: