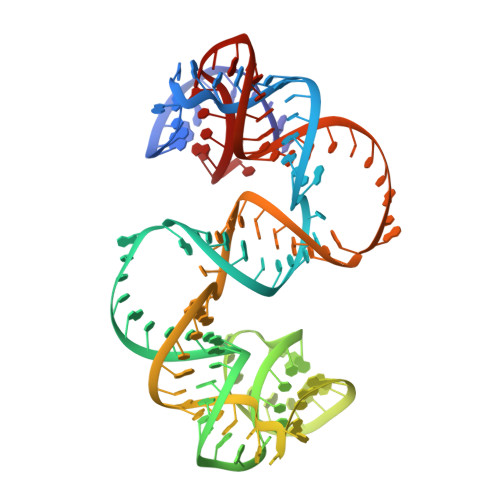
Structures of artificially designed discrete RNA nanoarchitectures at near-atomic resolution.
Liu, D., Shao, Y., Piccirilli, J.A., Weizmann, Y.(2021) Sci Adv 7: eabf4459-eabf4459
- PubMed: 34550747 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abf4459
- Primary Citation Related Structures:
7JRR, 7JRS, 7JRT - PubMed Abstract:
Although advances in nanotechnology have enabled the construction of complex and functional synthetic nucleic acid–based nanoarchitectures, high-resolution discrete structures are lacking because of the difficulty in obtaining good diffracting crystals. Here, we report the design and construction of RNA nanostructures based on homooligomerizable one-stranded tiles for x-ray crystallographic determination. We solved three structures to near-atomic resolution: a 2D parallelogram, a 3D nanobracelet unexpectedly formed from an RNA designed for a nanocage, and, eventually, a bona fide 3D nanocage designed with the guidance of the two previous structures. Structural details of their constituent motifs, such as kissing loops, branched kissing loops, and T-junctions, that resemble natural RNA motifs and resisted x-ray determination are revealed, providing insights into those natural motifs. This work unveils the largely unexplored potential of crystallography in gaining high-resolution feedback for nanoarchitectural design and suggests a route to investigate RNA motif structures by configuring them into nanoarchitectures.
- Department of Chemistry, The University of Chicago, Chicago, IL 60637, USA.
Organizational Affiliation: